| BQS OBSP Determination: |
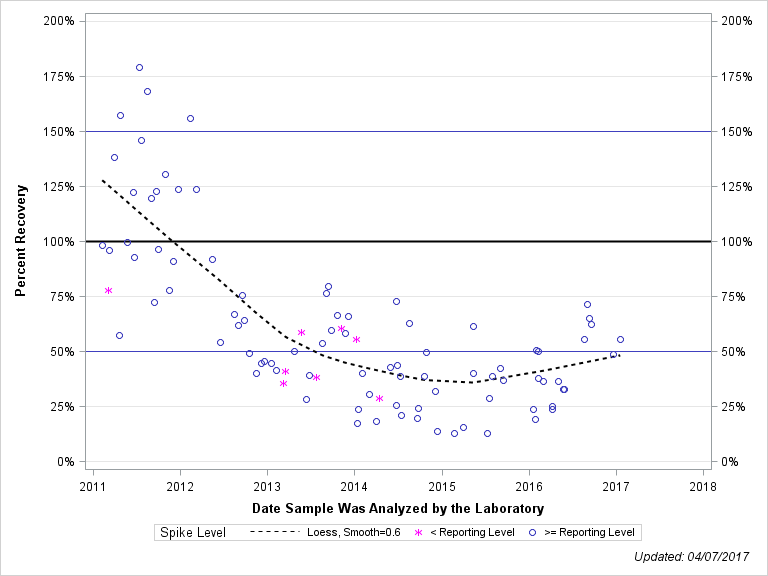
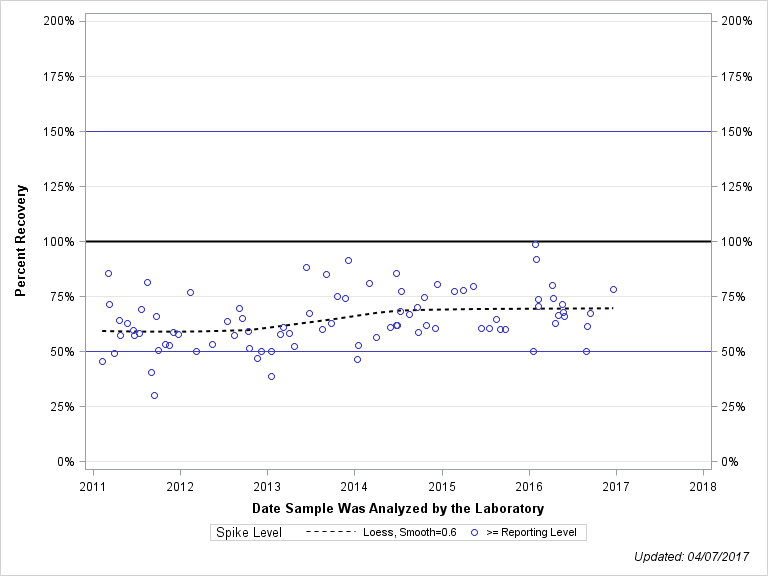
| 1-NAPHTHOL |
| METHOD: 2033 , TESTIDCODE: 49295GCM39 |
| Open Data Set |
| Plotted Points Statistics |
| 1-NAPHTHOL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 8 | 49% | 16% | 48% | 17% |
| >= Reporting Level | 87 | 62% | 39% | 50% | 30% |
| Total | 95 | 61% | 38% | 50% | 29% |
| Sample Statistics |
| 1-NAPHTHOL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 95 | 100% | Spiked |
| Estimated Values | 95 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 95 | . | |
| False Negatives | 0 | 0% | 0 out of 95 |
| Not Spiked | 74 | . | |
| False Positives | 0 | 0% | 0 out of 74 |
| Plotted Points Statistics |
| 2,6-DIETHYLANILINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 92% | 19% | 93% | 14% |
| Total | 94 | 92% | 19% | 93% | 14% |
| Sample Statistics |
| 2,6-DIETHYLANILINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| BQS OBSP Determination: |
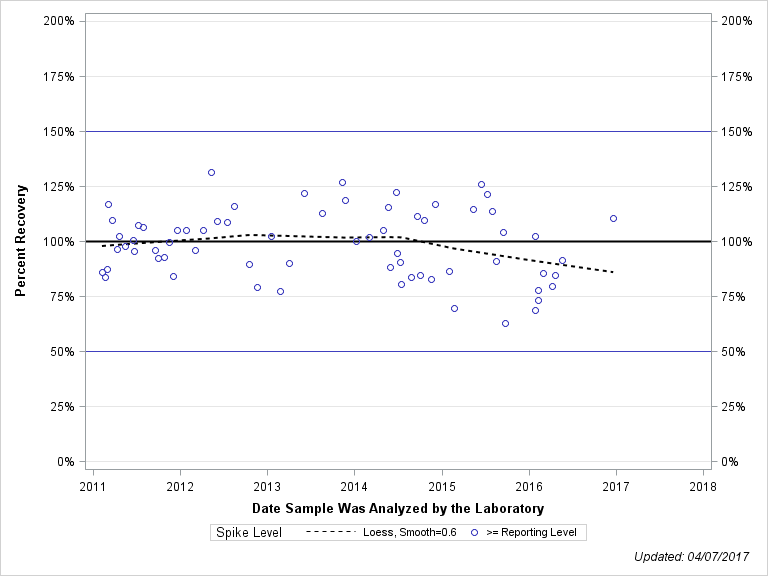
| 2-CHLORO-2,6-DIETHYLACETANILIDE |
| METHOD: 2033 , TESTIDCODE: 61618GCM39 |
| Open Data Set |
| Plotted Points Statistics |
| 2-CHLORO-2,6-DIETHYLACETANILIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 67 | 99% | 15% | 100% | 17% |
| Total | 67 | 99% | 15% | 100% | 17% |
| Sample Statistics |
| 2-CHLORO-2,6-DIETHYLACETANILIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 67 | 99% | Spiked |
| Estimated Values | 2 | 3% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 1% | Spiked |
| Spiked | 68 | . | |
| False Negatives | 0 | 0% | 0 out of 68 |
| Not Spiked | 101 | . | |
| False Positives | 1 | 1% | 1 out of 101 |
| Plotted Points Statistics |
| 2-ETHYL-6-METHYLANILINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 69 | 90% | 15% | 92% | 13% |
| Total | 69 | 90% | 15% | 92% | 13% |
| Sample Statistics |
| 2-ETHYL-6-METHYLANILINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 69 | 99% | Spiked |
| Estimated Values | 69 | 99% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 70 | . | |
| False Negatives | 1 | 1% | 1 out of 70 |
| Not Spiked | 92 | . | |
| False Positives | 0 | 0% | 0 out of 92 |
| Plotted Points Statistics |
| 3,4-DICHLOROANILINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 75% | 12% | 75% | 12% |
| Total | 94 | 75% | 12% | 75% | 12% |
| Sample Statistics |
| 3,4-DICHLOROANILINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 94 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| 3,5-DICHLOROANILINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 90 | 88% | 11% | 90% | 11% |
| Total | 90 | 88% | 11% | 90% | 11% |
| Sample Statistics |
| 3,5-DICHLOROANILINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 90 | 96% | Spiked |
| Estimated Values | 2 | 2% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 1% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 3 | 3% | 3 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| 4-CHLORO-2-METHYLPHENOL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 64% | 12% | 64% | 13% |
| Total | 94 | 64% | 12% | 64% | 13% |
| Sample Statistics |
| 4-CHLORO-2-METHYLPHENOL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 94 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| ACETOCHLOR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 95 | 89% | 16% | 88% | 16% |
| Total | 95 | 89% | 16% | 88% | 16% |
| Sample Statistics |
| ACETOCHLOR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 95 | 100% | Spiked |
| Estimated Values | 6 | 6% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 95 | . | |
| False Negatives | 0 | 0% | 0 out of 95 |
| Not Spiked | 74 | . | |
| False Positives | 1 | 1% | 1 out of 74 |
| Plotted Points Statistics |
| ALACHLOR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 92% | 14% | 90% | 13% |
| Total | 94 | 92% | 14% | 90% | 13% |
| Sample Statistics |
| ALACHLOR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 2 | 2% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| BQS OBSP Determination: |
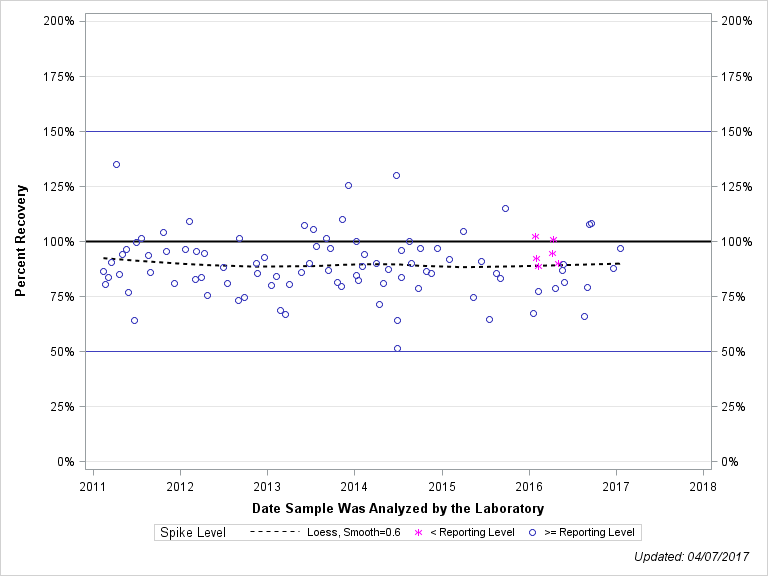
| ALPHA-ENDOSULFAN (ENDOSULFAN I) |
| METHOD: 2033 , TESTIDCODE: 34362GCM39 |
| Open Data Set |
| Plotted Points Statistics |
| ALPHA-ENDOSULFAN (ENDOSULFAN I) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 6 | 95% | 6% | 93% | 8% |
| >= Reporting Level | 90 | 89% | 14% | 87% | 12% |
| Total | 96 | 89% | 14% | 88% | 12% |
| Sample Statistics |
| ALPHA-ENDOSULFAN (ENDOSULFAN I) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 96 | 100% | Spiked |
| Estimated Values | 4 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 96 | . | |
| False Negatives | 0 | 0% | 0 out of 96 |
| Not Spiked | 73 | . | |
| False Positives | 0 | 0% | 0 out of 73 |
| Plotted Points Statistics |
| ATRAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 92% | 13% | 92% | 15% |
| Total | 94 | 92% | 13% | 92% | 15% |
| Sample Statistics |
| ATRAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 3 | 3% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| AZINPHOS-METHYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 7 | 92% | 19% | 91% | 11% |
| >= Reporting Level | 86 | 71% | 18% | 66% | 17% |
| Total | 93 | 73% | 19% | 71% | 20% |
| Sample Statistics |
| AZINPHOS-METHYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 93 | 100% | Spiked |
| Estimated Values | 93 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 0 | 0% | 0 out of 93 |
| Not Spiked | 76 | . | |
| False Positives | 0 | 0% | 0 out of 76 |
| Plotted Points Statistics |
| AZINPHOS-METHYL-OXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 45 | 71% | 25% | 65% | 16% |
| Total | 45 | 71% | 25% | 65% | 16% |
| Sample Statistics |
| AZINPHOS-METHYL-OXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 45 | 78% | Spiked |
| Estimated Values | 45 | 78% | Spiked |
| Deleted Values | 1 | 1% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 2% | Spiked |
| Spiked | 58 | . | |
| False Negatives | 12 | 21% | 12 out of 58 |
| Not Spiked | 111 | . | |
| False Positives | 1 | 1% | 1 out of 110 |
| Plotted Points Statistics |
| BENFLURALIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 87 | 77% | 14% | 77% | 15% |
| Total | 87 | 77% | 14% | 77% | 15% |
| Sample Statistics |
| BENFLURALIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 87 | 100% | Spiked |
| Estimated Values | 2 | 2% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 0 | 0% | 0 out of 87 |
| Not Spiked | 82 | . | |
| False Positives | 1 | 1% | 1 out of 82 |
| Plotted Points Statistics |
| CARBARYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 1 | 117% | . | 117% | 0% |
| >= Reporting Level | 94 | 78% | 18% | 79% | 19% |
| Total | 95 | 79% | 19% | 79% | 20% |
| Sample Statistics |
| CARBARYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 95 | 100% | Spiked |
| Estimated Values | 95 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 95 | . | |
| False Negatives | 0 | 0% | 0 out of 95 |
| Not Spiked | 74 | . | |
| False Positives | 3 | 4% | 3 out of 74 |
| Plotted Points Statistics |
| CARBOFURAN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 1 | 138% | . | 138% | 0% |
| >= Reporting Level | 92 | 89% | 21% | 91% | 23% |
| Total | 93 | 90% | 21% | 91% | 23% |
| Sample Statistics |
| CARBOFURAN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 93 | 100% | Spiked |
| Estimated Values | 93 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 0 | 0% | 0 out of 93 |
| Not Spiked | 76 | . | |
| False Positives | 0 | 0% | 0 out of 76 |
| Plotted Points Statistics |
| CHLORPYRIFOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 95 | 82% | 15% | 81% | 14% |
| Total | 95 | 82% | 15% | 81% | 14% |
| Sample Statistics |
| CHLORPYRIFOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 95 | 100% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 95 | . | |
| False Negatives | 0 | 0% | 0 out of 95 |
| Not Spiked | 74 | . | |
| False Positives | 0 | 0% | 0 out of 74 |
| BQS OBSP Determination: |
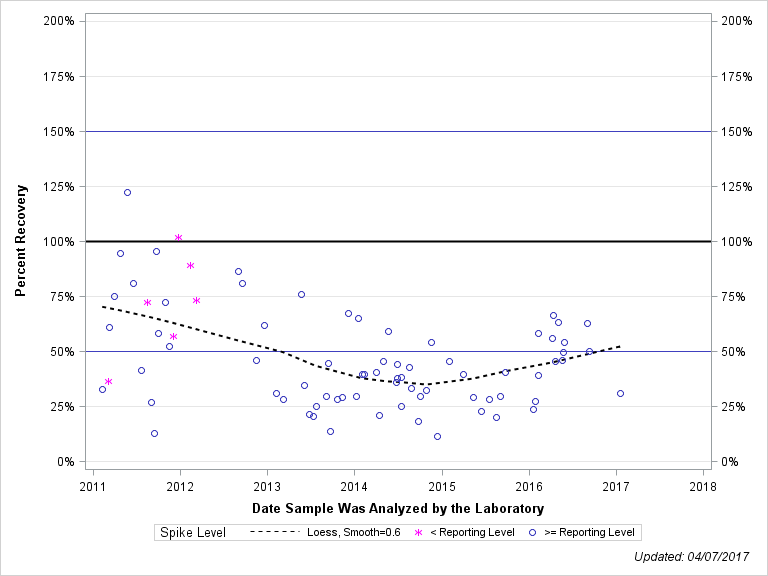
| CHLORPYRIFOS, OXYGEN ANALOG |
| METHOD: 2033 , TESTIDCODE: 61636GCM39 |
| Open Data Set |
| Plotted Points Statistics |
| CHLORPYRIFOS, OXYGEN ANALOG |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 6 | 72% | 23% | 73% | 24% |
| >= Reporting Level | 72 | 45% | 22% | 40% | 22% |
| Total | 78 | 47% | 23% | 41% | 23% |
| Sample Statistics |
| CHLORPYRIFOS, OXYGEN ANALOG |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 78 | 94% | Spiked |
| Estimated Values | 78 | 94% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 83 | . | |
| False Negatives | 5 | 6% | 5 out of 83 |
| Not Spiked | 76 | . | |
| False Positives | 1 | 1% | 1 out of 76 |
| Plotted Points Statistics |
| CIS-PERMETHRIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 89 | 68% | 18% | 67% | 18% |
| Total | 89 | 68% | 18% | 67% | 18% |
| Sample Statistics |
| CIS-PERMETHRIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 89 | 100% | Spiked |
| Estimated Values | 4 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 89 | . | |
| False Negatives | 0 | 0% | 0 out of 89 |
| Not Spiked | 80 | . | |
| False Positives | 0 | 0% | 0 out of 80 |
| Plotted Points Statistics |
| CYANAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 88% | 23% | 89% | 21% |
| Total | 94 | 88% | 23% | 89% | 21% |
| Sample Statistics |
| CYANAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 3 | 3% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| CYFLUTHRIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 67 | 70% | 26% | 68% | 22% |
| Total | 67 | 70% | 26% | 68% | 22% |
| Sample Statistics |
| CYFLUTHRIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 67 | 89% | Spiked |
| Estimated Values | 67 | 89% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 75 | . | |
| False Negatives | 8 | 11% | 8 out of 75 |
| Not Spiked | 83 | . | |
| False Positives | 1 | 1% | 1 out of 83 |
| Plotted Points Statistics |
| CYHALOTHRIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 90 | 57% | 19% | 57% | 15% |
| Total | 90 | 57% | 19% | 57% | 15% |
| Sample Statistics |
| CYHALOTHRIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 90 | 97% | Spiked |
| Estimated Values | 90 | 97% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 3 | 3% | 3 out of 93 |
| Not Spiked | 76 | . | |
| False Positives | 0 | 0% | 0 out of 76 |
| Plotted Points Statistics |
| CYPERMETHRIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 93 | 68% | 18% | 65% | 14% |
| Total | 93 | 68% | 18% | 65% | 14% |
| Sample Statistics |
| CYPERMETHRIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 93 | 100% | Spiked |
| Estimated Values | 93 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 0 | 0% | 0 out of 93 |
| Not Spiked | 76 | . | |
| False Positives | 1 | 1% | 1 out of 76 |
| Plotted Points Statistics |
| DACTHAL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 99% | 12% | 98% | 10% |
| Total | 94 | 99% | 12% | 98% | 10% |
| Sample Statistics |
| DACTHAL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| DEETHYLATRAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 93 | 67% | 30% | 58% | 25% |
| Total | 93 | 67% | 30% | 58% | 25% |
| Sample Statistics |
| DEETHYLATRAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 93 | 100% | Spiked |
| Estimated Values | 93 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 0 | 0% | 0 out of 93 |
| Not Spiked | 76 | . | |
| False Positives | 1 | 1% | 1 out of 76 |
| Plotted Points Statistics |
| DESULFINYLFIPRONIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 55 | 79% | 15% | 76% | 15% |
| Total | 55 | 79% | 15% | 76% | 15% |
| Sample Statistics |
| DESULFINYLFIPRONIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 55 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 55 | . | |
| False Negatives | 0 | 0% | 0 out of 55 |
| Not Spiked | 114 | . | |
| False Positives | 1 | 1% | 1 out of 114 |
| BQS OBSP Determination: |
| DESULFINYLFIPRONIL AMIDE |
| METHOD: 2033 , TESTIDCODE: 62169GCM29 |
| Open Data Set |
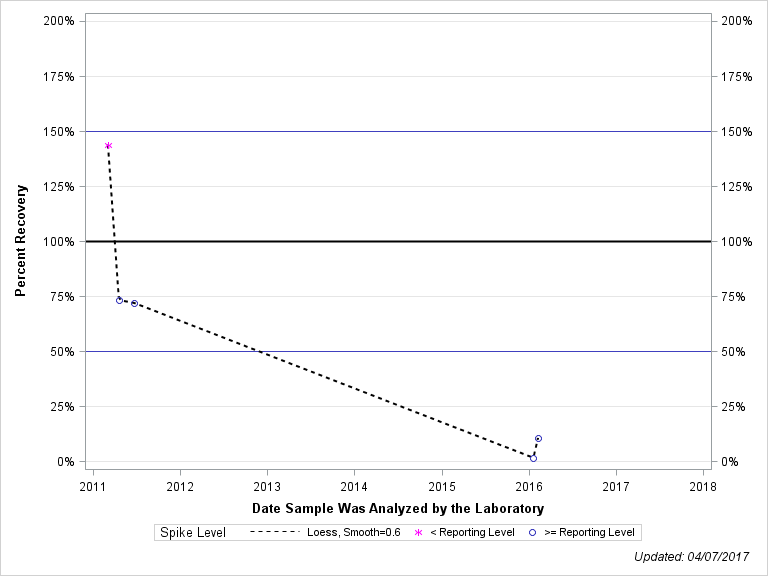
| Plotted Points Statistics |
| DESULFINYLFIPRONIL AMIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 1 | 144% | . | 144% | 0% |
| >= Reporting Level | 4 | 39% | 39% | 41% | 49% |
| Total | 5 | 60% | 57% | 72% | 46% |
| Sample Statistics |
| DESULFINYLFIPRONIL AMIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 5 | 50% | Spiked |
| Estimated Values | 5 | 50% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 10 | . | |
| False Negatives | 5 | 50% | 5 out of 10 |
| Not Spiked | 159 | . | |
| False Positives | 0 | 0% | 0 out of 159 |
| Plotted Points Statistics |
| DIAZINON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 96 | 96% | 12% | 97% | 11% |
| Total | 96 | 96% | 12% | 97% | 11% |
| Sample Statistics |
| DIAZINON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 96 | 100% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 96 | . | |
| False Negatives | 0 | 0% | 0 out of 96 |
| Not Spiked | 73 | . | |
| False Positives | 3 | 4% | 3 out of 73 |
| Plotted Points Statistics |
| DIAZINON, OXYGEN ANALOG |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 84 | 99% | 27% | 92% | 24% |
| Total | 84 | 99% | 27% | 92% | 24% |
| Sample Statistics |
| DIAZINON, OXYGEN ANALOG |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 84 | 97% | Spiked |
| Estimated Values | 9 | 10% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 1% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 2 | 2% | 2 out of 87 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| BQS OBSP Determination: |
| DICHLOROVOS (DICHLORVOS) |
| METHOD: 2033 , TESTIDCODE: 38775GCM39 |
| Open Data Set |
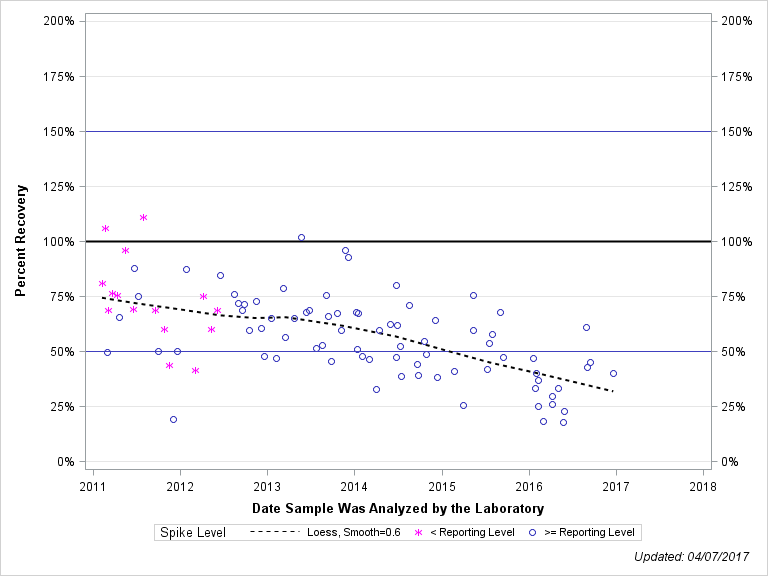
| Plotted Points Statistics |
| DICHLOROVOS (DICHLORVOS) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 15 | 73% | 20% | 69% | 15% |
| >= Reporting Level | 78 | 55% | 19% | 54% | 18% |
| Total | 93 | 58% | 20% | 60% | 18% |
| Sample Statistics |
| DICHLOROVOS (DICHLORVOS) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 93 | 100% | Spiked |
| Estimated Values | 93 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 0 | 0% | 0 out of 93 |
| Not Spiked | 76 | . | |
| False Positives | 1 | 1% | 1 out of 76 |
| Plotted Points Statistics |
| DICROTOPHOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 2 | 42% | 3% | 42% | 3% |
| >= Reporting Level | 82 | 26% | 10% | 25% | 9% |
| Total | 84 | 27% | 10% | 25% | 10% |
| Sample Statistics |
| DICROTOPHOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 84 | 97% | Spiked |
| Estimated Values | 84 | 97% | Spiked |
| Deleted Values | 2 | 1% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 3 | 3% | 3 out of 87 |
| Not Spiked | 74 | . | |
| False Positives | 1 | 1% | 1 out of 72 |
| Plotted Points Statistics |
| DIELDRIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 96 | 88% | 15% | 87% | 12% |
| Total | 96 | 88% | 15% | 87% | 12% |
| Sample Statistics |
| DIELDRIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 96 | 100% | Spiked |
| Estimated Values | 5 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 96 | . | |
| False Negatives | 0 | 0% | 0 out of 96 |
| Not Spiked | 73 | . | |
| False Positives | 1 | 1% | 1 out of 73 |
| Plotted Points Statistics |
| DIMETHOATE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 85 | 45% | 13% | 46% | 12% |
| Total | 85 | 45% | 13% | 46% | 12% |
| Sample Statistics |
| DIMETHOATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 85 | 98% | Spiked |
| Estimated Values | 85 | 98% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 2 | 2% | 2 out of 87 |
| Not Spiked | 82 | . | |
| False Positives | 0 | 0% | 0 out of 82 |
| Plotted Points Statistics |
| DISULFOTON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 1 | 77% | . | 77% | 0% |
| >= Reporting Level | 81 | 47% | 21% | 50% | 24% |
| Total | 82 | 47% | 21% | 51% | 24% |
| Sample Statistics |
| DISULFOTON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 82 | 90% | Spiked |
| Estimated Values | 82 | 90% | Spiked |
| Deleted Values | 5 | 3% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 95 | . | |
| False Negatives | 9 | 10% | 9 out of 91 |
| Not Spiked | 74 | . | |
| False Positives | 0 | 0% | 0 out of 73 |
| Plotted Points Statistics |
| DISULFOTON SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 74 | 99% | 35% | 91% | 16% |
| Total | 74 | 99% | 35% | 91% | 16% |
| Sample Statistics |
| DISULFOTON SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 74 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 74 | . | |
| False Negatives | 0 | 0% | 0 out of 74 |
| Not Spiked | 89 | . | |
| False Positives | 4 | 4% | 4 out of 89 |
| Plotted Points Statistics |
| ENDOSULFAN SULFATE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 88 | 81% | 13% | 80% | 13% |
| Total | 88 | 81% | 13% | 80% | 13% |
| Sample Statistics |
| ENDOSULFAN SULFATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 88 | 99% | Spiked |
| Estimated Values | 2 | 2% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 89 | . | |
| False Negatives | 1 | 1% | 1 out of 89 |
| Not Spiked | 80 | . | |
| False Positives | 0 | 0% | 0 out of 80 |
| Plotted Points Statistics |
| EPTC |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 97% | 11% | 97% | 9% |
| Total | 94 | 97% | 11% | 97% | 9% |
| Sample Statistics |
| EPTC |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 3 | 3% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| ETHION |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 86 | 97% | 22% | 94% | 20% |
| Total | 86 | 97% | 22% | 94% | 20% |
| Sample Statistics |
| ETHION |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 86 | 100% | Spiked |
| Estimated Values | 2 | 2% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 86 | . | |
| False Negatives | 0 | 0% | 0 out of 86 |
| Not Spiked | 83 | . | |
| False Positives | 2 | 2% | 2 out of 83 |
| Plotted Points Statistics |
| ETHION MONOXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 3 | 91% | 22% | 82% | 31% |
| Total | 3 | 91% | 22% | 82% | 31% |
| Sample Statistics |
| ETHION MONOXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 3 | 100% | Spiked |
| Estimated Values | 3 | 100% | Spiked |
| Deleted Values | 7 | 4% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 3 | . | |
| False Negatives | 0 | 0% | 0 out of 3 |
| Not Spiked | 166 | . | |
| False Positives | 2 | 1% | 2 out of 159 |
| Plotted Points Statistics |
| ETHOPROPHOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 77 | 93% | 16% | 93% | 19% |
| Total | 77 | 93% | 16% | 93% | 19% |
| Sample Statistics |
| ETHOPROPHOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 77 | 89% | Spiked |
| Estimated Values | 11 | 13% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 3 | 3% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 7 | 8% | 7 out of 87 |
| Not Spiked | 82 | . | |
| False Positives | 2 | 2% | 2 out of 82 |
| Plotted Points Statistics |
| FENAMIPHOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 1 | 114% | . | 114% | 0% |
| >= Reporting Level | 86 | 78% | 24% | 77% | 17% |
| Total | 87 | 78% | 24% | 78% | 17% |
| Sample Statistics |
| FENAMIPHOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 87 | 97% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 4 | 2% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 3 | 3% | 3 out of 90 |
| Not Spiked | 76 | . | |
| False Positives | 1 | 1% | 1 out of 75 |
| Plotted Points Statistics |
| FENAMIPHOS SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 1 | 196% | . | 196% | 0% |
| >= Reporting Level | 92 | 83% | 24% | 78% | 17% |
| Total | 93 | 84% | 27% | 79% | 17% |
| Sample Statistics |
| FENAMIPHOS SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 93 | 100% | Spiked |
| Estimated Values | 14 | 15% | Spiked |
| Deleted Values | 1 | 1% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 0 | 0% | 0 out of 93 |
| Not Spiked | 76 | . | |
| False Positives | 2 | 3% | 2 out of 75 |
| Plotted Points Statistics |
| FENAMIPHOS SULFOXIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 1 | 11% | . | 11% | 0% |
| >= Reporting Level | 74 | 30% | 24% | 24% | 15% |
| Total | 75 | 30% | 24% | 23% | 15% |
| Sample Statistics |
| FENAMIPHOS SULFOXIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 75 | 90% | Spiked |
| Estimated Values | 75 | 90% | Spiked |
| Deleted Values | 1 | 1% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 1% | Spiked |
| Spiked | 84 | . | |
| False Negatives | 7 | 8% | 7 out of 83 |
| Not Spiked | 65 | . | |
| False Positives | 0 | 0% | 0 out of 65 |
| Plotted Points Statistics |
| FIPRONIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 85 | 103% | 20% | 103% | 19% |
| Total | 85 | 103% | 20% | 103% | 19% |
| Sample Statistics |
| FIPRONIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 85 | 98% | Spiked |
| Estimated Values | 85 | 98% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 2 | 2% | 2 out of 87 |
| Not Spiked | 82 | . | |
| False Positives | 3 | 4% | 3 out of 82 |
| Plotted Points Statistics |
| FIPRONIL SULFIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 86 | 95% | 17% | 93% | 13% |
| Total | 86 | 95% | 17% | 93% | 13% |
| Sample Statistics |
| FIPRONIL SULFIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 86 | 99% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 1 | 1% | 1 out of 87 |
| Not Spiked | 62 | . | |
| False Positives | 3 | 5% | 3 out of 62 |
| Plotted Points Statistics |
| FIPRONIL SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 79 | 86% | 21% | 81% | 16% |
| Total | 79 | 86% | 21% | 81% | 16% |
| Sample Statistics |
| FIPRONIL SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 79 | 99% | Spiked |
| Estimated Values | 6 | 8% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 80 | . | |
| False Negatives | 1 | 1% | 1 out of 80 |
| Not Spiked | 75 | . | |
| False Positives | 2 | 3% | 2 out of 75 |
| Plotted Points Statistics |
| FONOFOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 87 | 85% | 18% | 84% | 12% |
| Total | 87 | 85% | 18% | 84% | 12% |
| Sample Statistics |
| FONOFOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 87 | 99% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 88 | . | |
| False Negatives | 1 | 1% | 1 out of 88 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| HEXAZINONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 89 | 63% | 20% | 61% | 21% |
| Total | 89 | 63% | 20% | 61% | 21% |
| Sample Statistics |
| HEXAZINONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 89 | 99% | Spiked |
| Estimated Values | 8 | 9% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 90 | . | |
| False Negatives | 1 | 1% | 1 out of 90 |
| Not Spiked | 79 | . | |
| False Positives | 0 | 0% | 0 out of 79 |
| Plotted Points Statistics |
| IPRODIONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 80 | 84% | 20% | 85% | 16% |
| Total | 80 | 84% | 20% | 85% | 16% |
| Sample Statistics |
| IPRODIONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 80 | 99% | Spiked |
| Estimated Values | 80 | 99% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 81 | . | |
| False Negatives | 1 | 1% | 1 out of 81 |
| Not Spiked | 80 | . | |
| False Positives | 0 | 0% | 0 out of 80 |
| Plotted Points Statistics |
| ISOFENPHOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 85 | 94% | 26% | 95% | 21% |
| Total | 85 | 94% | 26% | 95% | 21% |
| Sample Statistics |
| ISOFENPHOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 85 | 99% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 1 | 1% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 1 | 1% | 1 out of 86 |
| Not Spiked | 82 | . | |
| False Positives | 0 | 0% | 0 out of 82 |
| Plotted Points Statistics |
| MALAOXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 93 | 97% | 22% | 95% | 23% |
| Total | 93 | 97% | 22% | 95% | 23% |
| Sample Statistics |
| MALAOXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 93 | 100% | Spiked |
| Estimated Values | 8 | 9% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 0 | 0% | 0 out of 93 |
| Not Spiked | 76 | . | |
| False Positives | 3 | 4% | 3 out of 76 |
| Plotted Points Statistics |
| MALATHION |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 86 | 90% | 18% | 89% | 16% |
| Total | 86 | 90% | 18% | 89% | 16% |
| Sample Statistics |
| MALATHION |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 86 | 97% | Spiked |
| Estimated Values | 2 | 2% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 89 | . | |
| False Negatives | 3 | 3% | 3 out of 89 |
| Not Spiked | 80 | . | |
| False Positives | 0 | 0% | 0 out of 80 |
| Plotted Points Statistics |
| METALAXYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 87 | 94% | 17% | 93% | 16% |
| Total | 87 | 94% | 17% | 93% | 16% |
| Sample Statistics |
| METALAXYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 87 | 99% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 88 | . | |
| False Negatives | 1 | 1% | 1 out of 88 |
| Not Spiked | 81 | . | |
| False Positives | 0 | 0% | 0 out of 81 |
| BQS OBSP Determination: |
| METHIDATHION (SUPRACIDE) |
| METHOD: 2033 , TESTIDCODE: 61598GCM39 |
| Open Data Set |
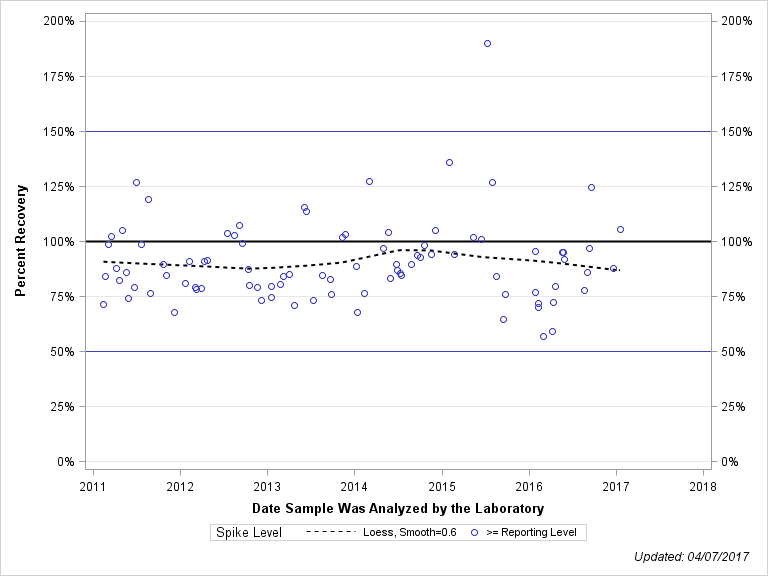
| Plotted Points Statistics |
| METHIDATHION (SUPRACIDE) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 89 | 91% | 19% | 87% | 14% |
| Total | 89 | 91% | 19% | 87% | 14% |
| Sample Statistics |
| METHIDATHION (SUPRACIDE) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 89 | 99% | Spiked |
| Estimated Values | 6 | 7% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 90 | . | |
| False Negatives | 1 | 1% | 1 out of 90 |
| Not Spiked | 79 | . | |
| False Positives | 0 | 0% | 0 out of 79 |
| Plotted Points Statistics |
| METOLACHLOR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 96 | 85% | 13% | 85% | 13% |
| Total | 96 | 85% | 13% | 85% | 13% |
| Sample Statistics |
| METOLACHLOR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 96 | 100% | Spiked |
| Estimated Values | 5 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 96 | . | |
| False Negatives | 0 | 0% | 0 out of 96 |
| Not Spiked | 73 | . | |
| False Positives | 0 | 0% | 0 out of 73 |
| Plotted Points Statistics |
| METRIBUZIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 89 | 82% | 19% | 81% | 17% |
| Total | 89 | 82% | 19% | 81% | 17% |
| Sample Statistics |
| METRIBUZIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 89 | 100% | Spiked |
| Estimated Values | 3 | 3% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 89 | . | |
| False Negatives | 0 | 0% | 0 out of 89 |
| Not Spiked | 80 | . | |
| False Positives | 0 | 0% | 0 out of 80 |
| Plotted Points Statistics |
| MOLINATE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 89 | 97% | 13% | 98% | 13% |
| Total | 89 | 97% | 13% | 98% | 13% |
| Sample Statistics |
| MOLINATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 89 | 95% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 2% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 3 | 3% | 3 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| MYCLOBUTANIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 78% | 14% | 77% | 11% |
| Total | 94 | 78% | 14% | 77% | 11% |
| Sample Statistics |
| MYCLOBUTANIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| OXYFLUORFEN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 85% | 19% | 84% | 16% |
| Total | 94 | 85% | 19% | 84% | 16% |
| Sample Statistics |
| OXYFLUORFEN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 4 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| PARAOXON-METHYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 87 | 93% | 22% | 92% | 22% |
| Total | 87 | 93% | 22% | 92% | 22% |
| Sample Statistics |
| PARAOXON-METHYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 87 | 97% | Spiked |
| Estimated Values | 87 | 97% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 90 | . | |
| False Negatives | 3 | 3% | 3 out of 90 |
| Not Spiked | 79 | . | |
| False Positives | 0 | 0% | 0 out of 79 |
| Plotted Points Statistics |
| PARATHION-METHYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 89 | 107% | 20% | 103% | 17% |
| Total | 89 | 107% | 20% | 103% | 17% |
| Sample Statistics |
| PARATHION-METHYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 89 | 99% | Spiked |
| Estimated Values | 9 | 10% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 90 | . | |
| False Negatives | 1 | 1% | 1 out of 90 |
| Not Spiked | 71 | . | |
| False Positives | 0 | 0% | 0 out of 71 |
| Plotted Points Statistics |
| PENDIMETHALIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 87 | 94% | 19% | 90% | 16% |
| Total | 87 | 94% | 19% | 90% | 16% |
| Sample Statistics |
| PENDIMETHALIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 87 | 100% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 0 | 0% | 0 out of 87 |
| Not Spiked | 82 | . | |
| False Positives | 0 | 0% | 0 out of 82 |
| Plotted Points Statistics |
| PHORATE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 79 | 65% | 20% | 68% | 21% |
| Total | 79 | 65% | 20% | 68% | 21% |
| Sample Statistics |
| PHORATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 79 | 91% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 1 | 1% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 8 | 9% | 8 out of 87 |
| Not Spiked | 74 | . | |
| False Positives | 0 | 0% | 0 out of 73 |
| Plotted Points Statistics |
| PHORATE OXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 1 | 118% | . | 118% | 0% |
| >= Reporting Level | 55 | 65% | 20% | 64% | 21% |
| Total | 56 | 66% | 21% | 64% | 20% |
| Sample Statistics |
| PHORATE OXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 56 | 95% | Spiked |
| Estimated Values | 56 | 95% | Spiked |
| Deleted Values | 7 | 4% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 64 | . | |
| False Negatives | 3 | 5% | 3 out of 59 |
| Not Spiked | 105 | . | |
| False Positives | 0 | 0% | 0 out of 103 |
| Plotted Points Statistics |
| PHOSMET (IMIDAN) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 7 | 68% | 17% | 68% | 24% |
| >= Reporting Level | 82 | 61% | 15% | 57% | 17% |
| Total | 89 | 62% | 16% | 57% | 17% |
| Sample Statistics |
| PHOSMET (IMIDAN) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 89 | 97% | Spiked |
| Estimated Values | 89 | 97% | Spiked |
| Deleted Values | 2 | 1% | Spiked + Not Spiked |
| Spiked, Censored | 3 | 3% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 0 | 0% | 0 out of 92 |
| Not Spiked | 76 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| PHOSMET OXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 56 | 80% | 56% | 70% | 23% |
| Total | 56 | 80% | 56% | 70% | 23% |
| Sample Statistics |
| PHOSMET OXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 56 | 93% | Spiked |
| Estimated Values | 56 | 93% | Spiked |
| Deleted Values | 9 | 5% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 2% | Spiked |
| Spiked | 64 | . | |
| False Negatives | 3 | 5% | 3 out of 60 |
| Not Spiked | 105 | . | |
| False Positives | 3 | 3% | 3 out of 100 |
| Plotted Points Statistics |
| PROMETON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 87 | 83% | 16% | 83% | 16% |
| Total | 87 | 83% | 16% | 83% | 16% |
| Sample Statistics |
| PROMETON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 87 | 100% | Spiked |
| Estimated Values | 2 | 2% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 0 | 0% | 0 out of 87 |
| Not Spiked | 82 | . | |
| False Positives | 0 | 0% | 0 out of 82 |
| Plotted Points Statistics |
| PROMETRYN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 88 | 81% | 18% | 82% | 14% |
| Total | 88 | 81% | 18% | 82% | 14% |
| Sample Statistics |
| PROMETRYN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 88 | 100% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 88 | . | |
| False Negatives | 0 | 0% | 0 out of 88 |
| Not Spiked | 74 | . | |
| False Positives | 1 | 1% | 1 out of 74 |
| Plotted Points Statistics |
| PROPANIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 87 | 105% | 19% | 103% | 19% |
| Total | 87 | 105% | 19% | 103% | 19% |
| Sample Statistics |
| PROPANIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 87 | 100% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 0 | 0% | 0 out of 87 |
| Not Spiked | 82 | . | |
| False Positives | 0 | 0% | 0 out of 82 |
| Plotted Points Statistics |
| PROPARGITE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 89 | 80% | 34% | 76% | 28% |
| Total | 89 | 80% | 34% | 76% | 28% |
| Sample Statistics |
| PROPARGITE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 89 | 96% | Spiked |
| Estimated Values | 16 | 17% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 1% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 3 | 3% | 3 out of 93 |
| Not Spiked | 76 | . | |
| False Positives | 0 | 0% | 0 out of 76 |
| Plotted Points Statistics |
| PROPYZAMIDE (PRONAMIDE) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 87% | 15% | 87% | 15% |
| Total | 94 | 87% | 15% | 87% | 15% |
| Sample Statistics |
| PROPYZAMIDE (PRONAMIDE) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| SIMAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 94 | 91% | 13% | 92% | 13% |
| Total | 94 | 91% | 13% | 92% | 13% |
| Sample Statistics |
| SIMAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 94 | 100% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 94 | . | |
| False Negatives | 0 | 0% | 0 out of 94 |
| Not Spiked | 75 | . | |
| False Positives | 0 | 0% | 0 out of 75 |
| Plotted Points Statistics |
| TEBUCONAZOLE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 73 | 66% | 15% | 64% | 13% |
| Total | 73 | 66% | 15% | 64% | 13% |
| Sample Statistics |
| TEBUCONAZOLE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 73 | 100% | Spiked |
| Estimated Values | 4 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 73 | . | |
| False Negatives | 0 | 0% | 0 out of 73 |
| Not Spiked | 96 | . | |
| False Positives | 0 | 0% | 0 out of 96 |
| Plotted Points Statistics |
| TEBUTHIURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 1 | 211% | . | 211% | 0% |
| >= Reporting Level | 92 | 126% | 37% | 123% | 33% |
| Total | 93 | 127% | 38% | 123% | 33% |
| Sample Statistics |
| TEBUTHIURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 93 | 100% | Spiked |
| Estimated Values | 28 | 30% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 93 | . | |
| False Negatives | 0 | 0% | 0 out of 93 |
| Not Spiked | 76 | . | |
| False Positives | 0 | 0% | 0 out of 76 |
| Plotted Points Statistics |
| TEFLUTHRIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 87 | 60% | 11% | 58% | 11% |
| Total | 87 | 60% | 11% | 58% | 11% |
| Sample Statistics |
| TEFLUTHRIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 87 | 100% | Spiked |
| Estimated Values | 87 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 87 | . | |
| False Negatives | 0 | 0% | 0 out of 87 |
| Not Spiked | 82 | . | |
| False Positives | 1 | 1% | 1 out of 82 |
| Plotted Points Statistics |
| TERBUFOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 86 | 59% | 16% | 59% | 14% |
| Total | 86 | 59% | 16% | 59% | 14% |
| Sample Statistics |
| TERBUFOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 86 | 93% | Spiked |
| Estimated Values | 1 | 1% | Spiked |
| Deleted Values | 5 | 3% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 95 | . | |
| False Negatives | 6 | 7% | 6 out of 92 |
| Not Spiked | 74 | . | |
| False Positives | 1 | 1% | 1 out of 72 |
| BQS OBSP Determination: |
| TERBUFOS-O-ANALOGUE SULFONE |
| METHOD: 2033 , TESTIDCODE: 61674GCM39 |
| Open Data Set |
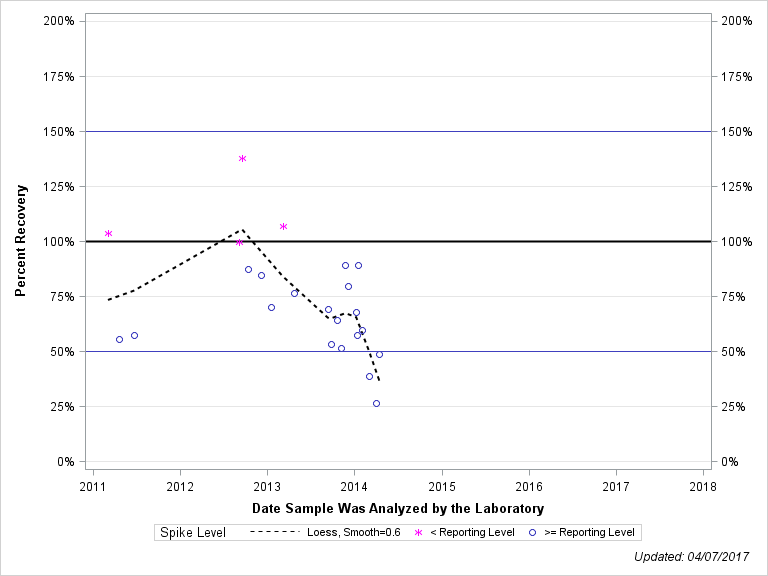
| Plotted Points Statistics |
| TERBUFOS-O-ANALOGUE SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 4 | 112% | 17% | 105% | 15% |
| >= Reporting Level | 19 | 64% | 17% | 64% | 19% |
| Total | 23 | 73% | 25% | 69% | 25% |
| Sample Statistics |
| TERBUFOS-O-ANALOGUE SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 23 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 23 | . | |
| False Negatives | 0 | 0% | 0 out of 23 |
| Not Spiked | 139 | . | |
| False Positives | 0 | 0% | 0 out of 139 |
| Plotted Points Statistics |
| TERBUTHYLAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 95 | 95% | 14% | 95% | 15% |
| Total | 95 | 95% | 14% | 95% | 15% |
| Sample Statistics |
| TERBUTHYLAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 95 | 99% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 1% | Spiked |
| Spiked | 96 | . | |
| False Negatives | 0 | 0% | 0 out of 96 |
| Not Spiked | 73 | . | |
| False Positives | 0 | 0% | 0 out of 73 |
| Plotted Points Statistics |
| THIOBENCARB |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 89 | 82% | 15% | 81% | 17% |
| Total | 89 | 82% | 15% | 81% | 17% |
| Sample Statistics |
| THIOBENCARB |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 89 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 89 | . | |
| False Negatives | 0 | 0% | 0 out of 89 |
| Not Spiked | 80 | . | |
| False Positives | 3 | 4% | 3 out of 80 |
| BQS OBSP Determination: |
| TRIBUFOS (S,S,S-TRIBUTYLPHOSPHOROTRITHIOATE (DEF)) |
| METHOD: 2033 , TESTIDCODE: 61610GCM39 |
| Open Data Set |
| Plotted Points Statistics |
| TRIBUFOS (S,S,S-TRIBUTYLPHOSPHOROTRITHIOATE (DEF)) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 86 | 64% | 13% | 62% | 12% |
| Total | 86 | 64% | 13% | 62% | 12% |
| Sample Statistics |
| TRIBUFOS (S,S,S-TRIBUTYLPHOSPHOROTRITHIOATE (DEF)) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 86 | 100% | Spiked |
| Estimated Values | 86 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 86 | . | |
| False Negatives | 0 | 0% | 0 out of 86 |
| Not Spiked | 83 | . | |
| False Positives | 2 | 2% | 2 out of 83 |