| BQS OBSP Determination: |
| 1H-1,2,4-TRIAZOLE |
| METHOD: 2437 , TESTIDCODE: 68498LCM60 |
| Open Data Set |
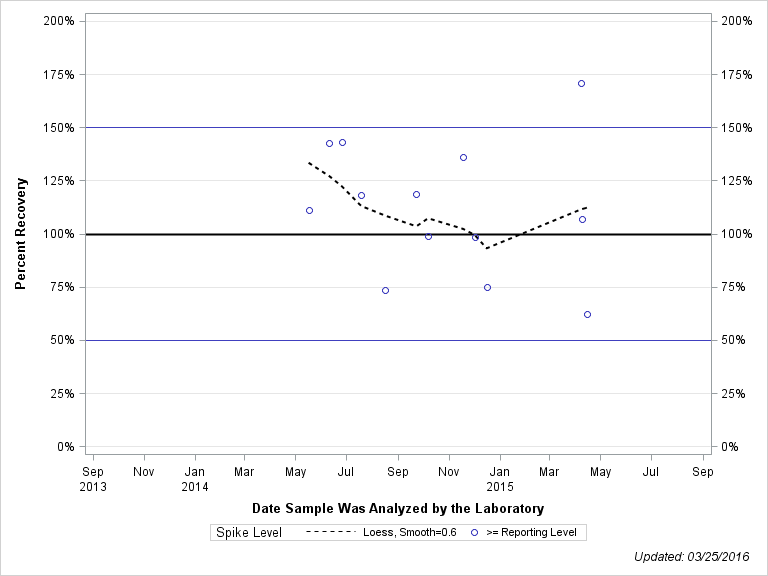
| Plotted Points Statistics |
| 1H-1,2,4-TRIAZOLE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 13 | 112% | 31% | 111% | 28% |
| Total | 13 | 112% | 31% | 111% | 28% |
| Sample Statistics |
| 1H-1,2,4-TRIAZOLE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 13 | 87% | Spiked |
| Estimated Values | 13 | 87% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 7% | Spiked |
| Spiked | 15 | . | |
| False Negatives | 1 | 7% | 1 out of 15 |
| Not Spiked | 33 | . | |
| False Positives | 0 | 0% | 0 out of 33 |
| Plotted Points Statistics |
| 2,4-D |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 123% | 53% | 112% | 36% |
| Total | 22 | 123% | 53% | 112% | 36% |
| Sample Statistics |
| 2,4-D |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 85% | Spiked |
| Estimated Values | 5 | 19% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 3 | 12% | 3 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| BQS OBSP Determination: |
| 2-(1-HYDROXYETHYL)-6-METHYLANILINE |
| METHOD: 2437 , TESTIDCODE: 68611LCM60 |
| Open Data Set |

| Sample Statistics |
| 2-(1-HYDROXYETHYL)-6-METHYLANILINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 1 | 2% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 47 |
| BQS OBSP Determination: |
| 2-AMINO-N-ISOPROPYLBENZAMIDE |
| METHOD: 2437 , TESTIDCODE: 68503LCM60 |
| Open Data Set |
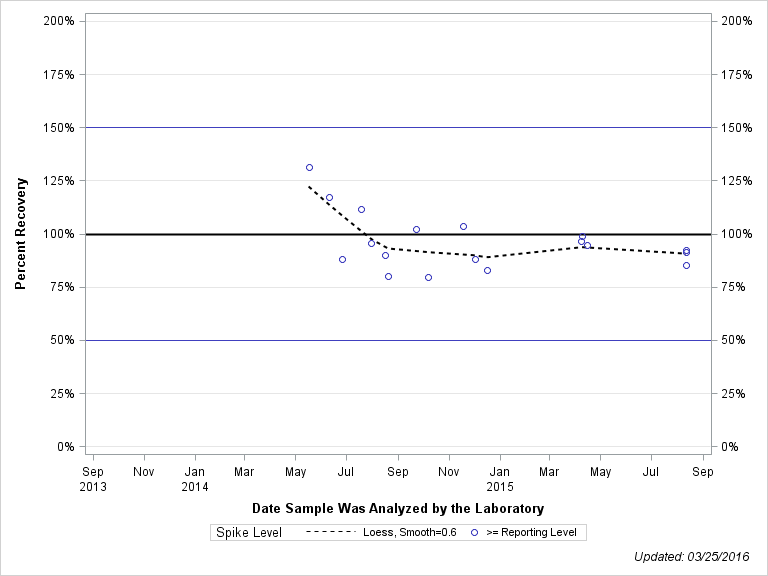
| Plotted Points Statistics |
| 2-AMINO-N-ISOPROPYLBENZAMIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 18 | 96% | 13% | 93% | 10% |
| Total | 18 | 96% | 13% | 93% | 10% |
| Sample Statistics |
| 2-AMINO-N-ISOPROPYLBENZAMIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 18 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 18 | . | |
| False Negatives | 0 | 0% | 0 out of 18 |
| Not Spiked | 30 | . | |
| False Positives | 1 | 3% | 1 out of 30 |
| Plotted Points Statistics |
| 2-AMINOBENZIMIDAZOLE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 18 | 120% | 17% | 114% | 16% |
| Total | 18 | 120% | 17% | 114% | 16% |
| Sample Statistics |
| 2-AMINOBENZIMIDAZOLE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 18 | 100% | Spiked |
| Estimated Values | 1 | 6% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 18 | . | |
| False Negatives | 0 | 0% | 0 out of 18 |
| Not Spiked | 30 | . | |
| False Positives | 0 | 0% | 0 out of 30 |
| BQS OBSP Determination: |
| 2-CHLORO-2',6'-DIETHYLACETANILIDE |
| METHOD: 2437 , TESTIDCODE: 68525LCM60 |
| Open Data Set |
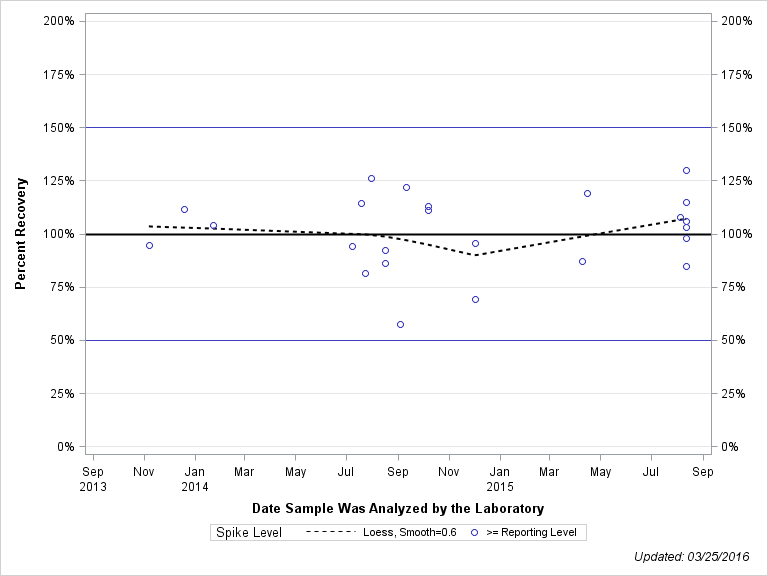
| Plotted Points Statistics |
| 2-CHLORO-2',6'-DIETHYLACETANILIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 24 | 101% | 18% | 104% | 18% |
| Total | 24 | 101% | 18% | 104% | 18% |
| Sample Statistics |
| 2-CHLORO-2',6'-DIETHYLACETANILIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 24 | 100% | Spiked |
| Estimated Values | 4 | 17% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 0 | 0% | 0 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 0 | 0% | 0 out of 24 |
| BQS OBSP Determination: |
| 2-CHLORO-N-(2-ETHYL-6-METHYLPHENYL)ACETAMIDE |
| METHOD: 2437 , TESTIDCODE: 68521LCM60 |
| Open Data Set |
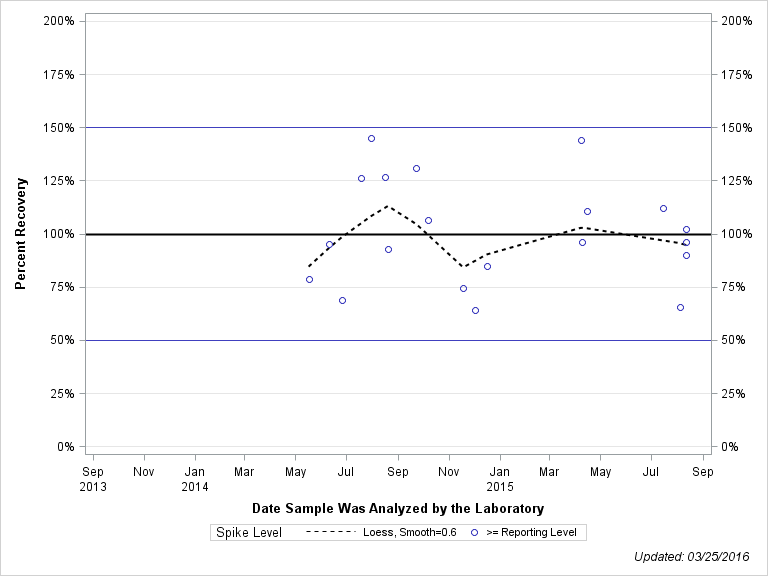
| Plotted Points Statistics |
| 2-CHLORO-N-(2-ETHYL-6-METHYLPHENYL)ACETAMIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 20 | 100% | 25% | 96% | 28% |
| Total | 20 | 100% | 25% | 96% | 28% |
| Sample Statistics |
| 2-CHLORO-N-(2-ETHYL-6-METHYLPHENYL)ACETAMIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 20 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 20 | . | |
| False Negatives | 0 | 0% | 0 out of 20 |
| Not Spiked | 28 | . | |
| False Positives | 3 | 11% | 3 out of 28 |
| BQS OBSP Determination: |
| 2-HYDROXY-4-ISOPROPYLAMINO-6-AMINO-S-TRIAZINE |
| METHOD: 2437 , TESTIDCODE: 68659LCM60 |
| Open Data Set |
| Plotted Points Statistics |
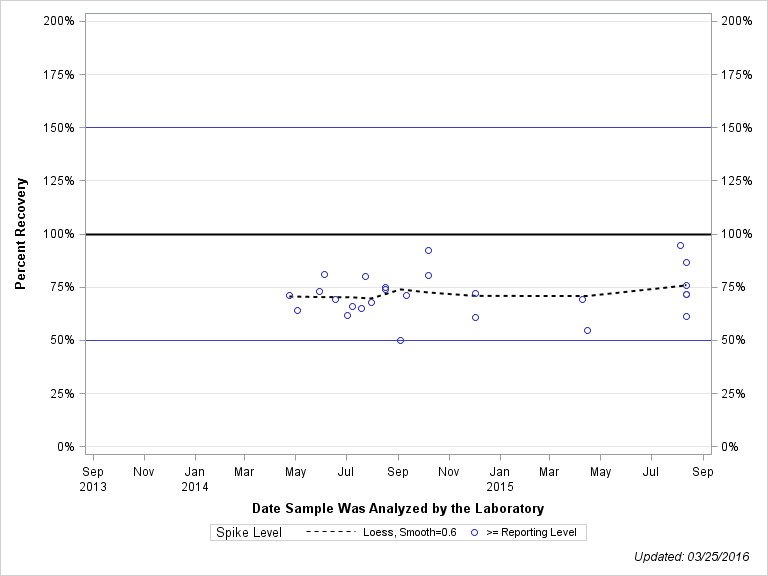
| 2-HYDROXY-4-ISOPROPYLAMINO-6-AMINO-S-TRIAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 72% | 10% | 71% | 8% |
| Total | 26 | 72% | 10% | 71% | 8% |
| Sample Statistics |
| 2-HYDROXY-4-ISOPROPYLAMINO-6-AMINO-S-TRIAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 96% | Spiked |
| Estimated Values | 2 | 7% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 1 | 4% | 1 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| BQS OBSP Determination: |
| 2-HYDROXY-4-ISOPROPYLAMINO-6-ETHYLAMINO-S-TRIAZINE (OIET) |
| METHOD: 2437 , TESTIDCODE: 68660LCM60 |
| Open Data Set |
| Plotted Points Statistics |
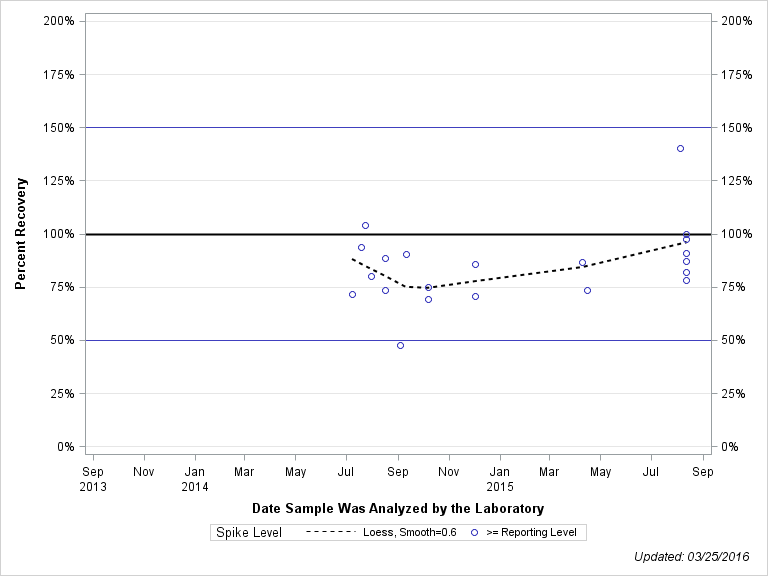
| 2-HYDROXY-4-ISOPROPYLAMINO-6-ETHYLAMINO-S-TRIAZINE (OIET) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 85% | 18% | 86% | 13% |
| Total | 21 | 85% | 18% | 86% | 13% |
| Sample Statistics |
| 2-HYDROXY-4-ISOPROPYLAMINO-6-ETHYLAMINO-S-TRIAZINE (OIET) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 100% | Spiked |
| Estimated Values | 7 | 33% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 21 | . | |
| False Negatives | 0 | 0% | 0 out of 21 |
| Not Spiked | 27 | . | |
| False Positives | 2 | 7% | 2 out of 27 |
| BQS OBSP Determination: |
| 2-HYDROXY-6-ETHYLAMINO-4-AMINO-S-TRIAZINE |
| METHOD: 2437 , TESTIDCODE: 68656LCM60 |
| Open Data Set |
| Plotted Points Statistics |
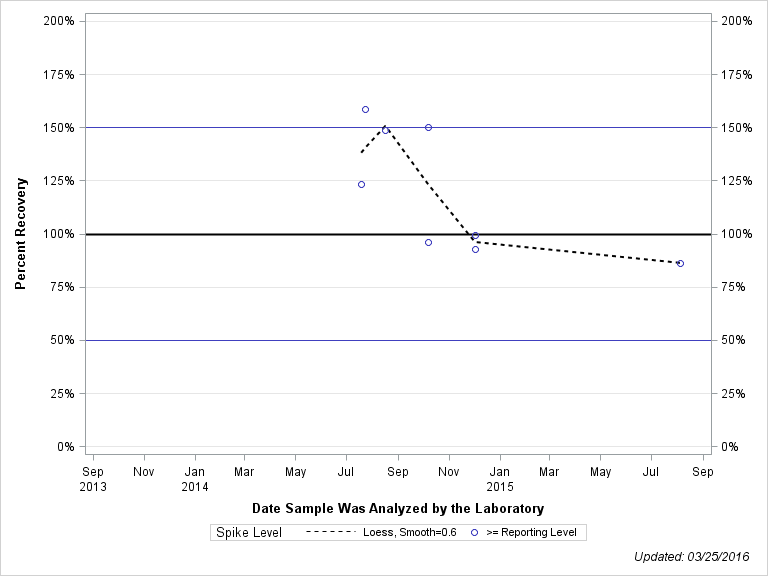
| 2-HYDROXY-6-ETHYLAMINO-4-AMINO-S-TRIAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 8 | 119% | 30% | 111% | 41% |
| Total | 8 | 119% | 30% | 111% | 41% |
| Sample Statistics |
| 2-HYDROXY-6-ETHYLAMINO-4-AMINO-S-TRIAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 8 | 38% | Spiked |
| Estimated Values | 2 | 10% | Spiked |
| Deleted Values | 2 | 4% | Spiked + Not Spiked |
| Spiked, Censored | 8 | 38% | Spiked |
| Spiked | 21 | . | |
| False Negatives | 5 | 24% | 5 out of 21 |
| Not Spiked | 27 | . | |
| False Positives | 1 | 4% | 1 out of 25 |
| BQS OBSP Determination: |
| 2-ISOPROPYL-6-METHYL-4-PYRIMIDINOL |
| METHOD: 2437 , TESTIDCODE: 68505LCM60 |
| Open Data Set |
| Plotted Points Statistics |
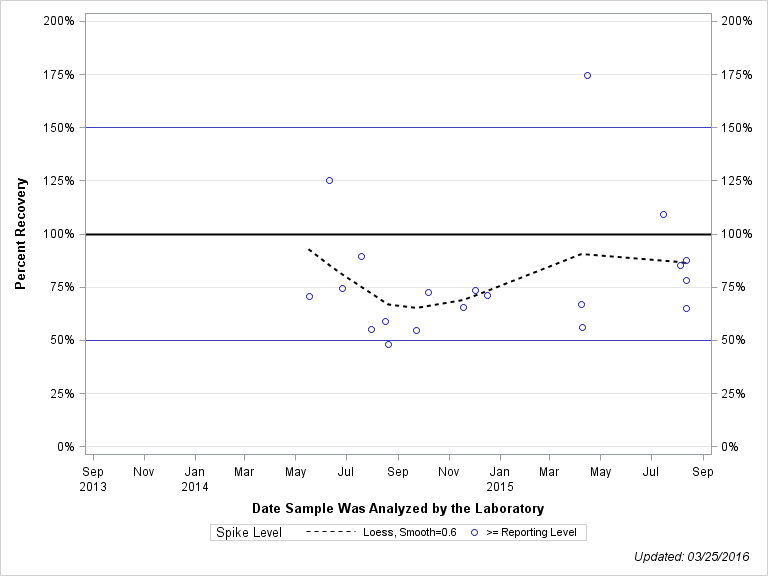
| 2-ISOPROPYL-6-METHYL-4-PYRIMIDINOL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 20 | 79% | 29% | 72% | 18% |
| Total | 20 | 79% | 29% | 72% | 18% |
| Sample Statistics |
| 2-ISOPROPYL-6-METHYL-4-PYRIMIDINOL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 20 | 100% | Spiked |
| Estimated Values | 15 | 75% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 20 | . | |
| False Negatives | 0 | 0% | 0 out of 20 |
| Not Spiked | 28 | . | |
| False Positives | 0 | 0% | 0 out of 28 |
| BQS OBSP Determination: |
| 2-[(2-ETHYL-6-METHYLPHENYL)AMINO]-1-PROPANOL |
| METHOD: 2437 , TESTIDCODE: 68595LCM60 |
| Open Data Set |
| Plotted Points Statistics |
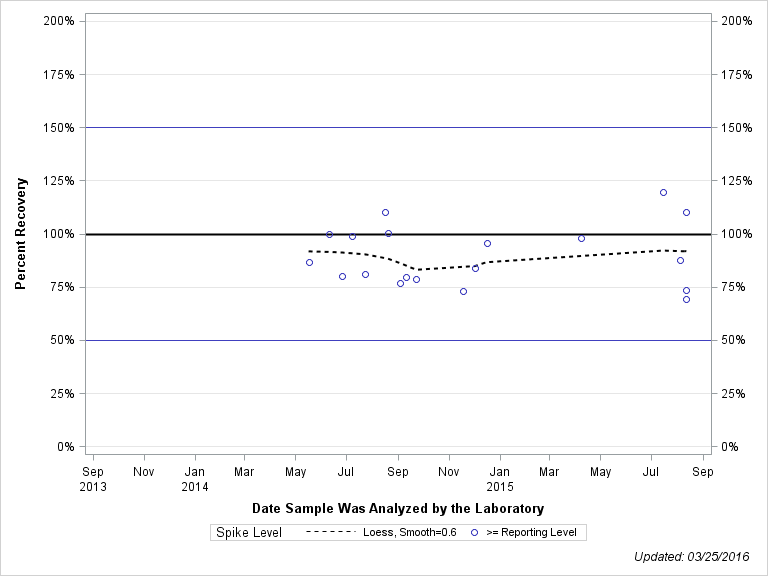
| 2-[(2-ETHYL-6-METHYLPHENYL)AMINO]-1-PROPANOL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 19 | 90% | 14% | 87% | 16% |
| Total | 19 | 90% | 14% | 87% | 16% |
| Sample Statistics |
| 2-[(2-ETHYL-6-METHYLPHENYL)AMINO]-1-PROPANOL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 19 | 95% | Spiked |
| Estimated Values | 1 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 20 | . | |
| False Negatives | 1 | 5% | 1 out of 20 |
| Not Spiked | 28 | . | |
| False Positives | 0 | 0% | 0 out of 28 |
| Plotted Points Statistics |
| 3,4-DICHLOROPHENYLUREA |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 7 | 104% | 11% | 106% | 15% |
| Total | 7 | 104% | 11% | 106% | 15% |
| Sample Statistics |
| 3,4-DICHLOROPHENYLUREA |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 7 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 7 | . | |
| False Negatives | 0 | 0% | 0 out of 7 |
| Not Spiked | 41 | . | |
| False Positives | 1 | 2% | 1 out of 41 |
| Plotted Points Statistics |
| 3-HYDROXYCARBOFURAN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 90% | 18% | 87% | 24% |
| Total | 26 | 90% | 18% | 87% | 24% |
| Sample Statistics |
| 3-HYDROXYCARBOFURAN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 100% | Spiked |
| Estimated Values | 26 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 0 | 0% | 0 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| 3-PHENOXYBENZOIC ACID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 15 | 100% | 18% | 98% | 14% |
| Total | 15 | 100% | 18% | 98% | 14% |
| Sample Statistics |
| 3-PHENOXYBENZOIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 15 | 100% | Spiked |
| Estimated Values | 1 | 7% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 15 | . | |
| False Negatives | 0 | 0% | 0 out of 15 |
| Not Spiked | 33 | . | |
| False Positives | 4 | 12% | 4 out of 33 |
| BQS OBSP Determination: |
| 4-(HYDROXYMETHYL)PENDIMETHALIN |
| METHOD: 2437 , TESTIDCODE: 68511LCM60 |
| Open Data Set |

| Sample Statistics |
| 4-(HYDROXYMETHYL)PENDIMETHALIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| BQS OBSP Determination: |
| 4-CHLOROBENZYLMETHYL SULFOXIDE |
| METHOD: 2437 , TESTIDCODE: 68514LCM60 |
| Open Data Set |

| Sample Statistics |
| 4-CHLOROBENZYLMETHYL SULFOXIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | 0% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 13 | . | |
| False Negatives | 13 | 100% | 13 out of 13 |
| Not Spiked | 35 | . | |
| False Positives | 0 | 0% | 0 out of 35 |
| Plotted Points Statistics |
| 4-HYDROXYCHLOROTHALONIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 3 | 110% | 21% | 110% | 31% |
| Total | 3 | 110% | 21% | 110% | 31% |
| Sample Statistics |
| 4-HYDROXYCHLOROTHALONIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 3 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 3 | . | |
| False Negatives | 0 | 0% | 0 out of 3 |
| Not Spiked | 45 | . | |
| False Positives | 0 | 0% | 0 out of 45 |
| Sample Statistics |
| 4-HYDROXYHEXAZINONE A |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Sample Statistics |
| 4-HYDROXY MOLINATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 1 | 2% | 1 out of 48 |
| Plotted Points Statistics |
| ACEPHATE (ORTHENE) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 24 | 76% | 17% | 73% | 9% |
| Total | 24 | 76% | 17% | 73% | 9% |
| Sample Statistics |
| ACEPHATE (ORTHENE) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 24 | 100% | Spiked |
| Estimated Values | 4 | 17% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 0 | 0% | 0 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 0 | 0% | 0 out of 24 |
| Plotted Points Statistics |
| ACETOCHLOR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 28 | 80% | 14% | 77% | 14% |
| Total | 28 | 80% | 14% | 77% | 14% |
| Sample Statistics |
| ACETOCHLOR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 28 | 100% | Spiked |
| Estimated Values | 2 | 7% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 0 | 0% | 0 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 1 | 5% | 1 out of 20 |
| BQS OBSP Determination: |
| ACETOCHLOR OXANILIC ACID |
| METHOD: 2437 , TESTIDCODE: 68522LCM60 |
| Open Data Set |
| Plotted Points Statistics |
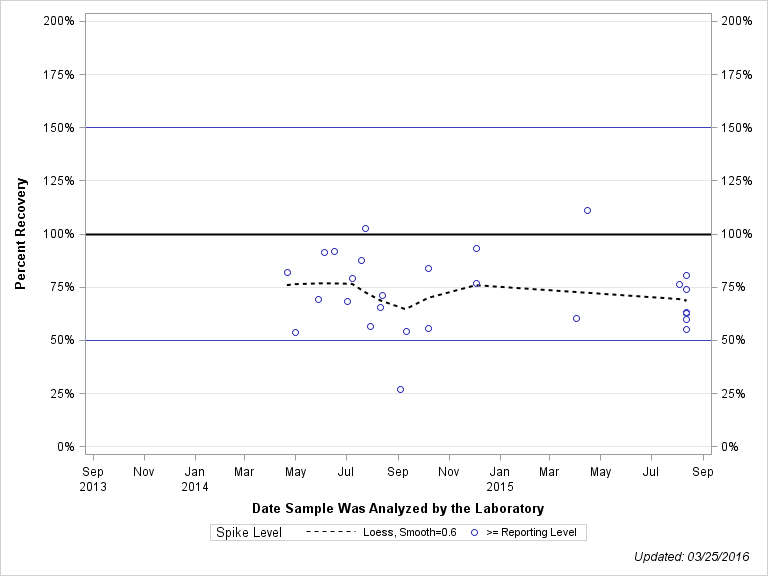
| ACETOCHLOR OXANILIC ACID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 27 | 72% | 18% | 71% | 18% |
| Total | 27 | 72% | 18% | 71% | 18% |
| Sample Statistics |
| ACETOCHLOR OXANILIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 27 | 100% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 0 | 0% | 0 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| BQS OBSP Determination: |
| ACETOCHLOR SULFONIC ACID |
| METHOD: 2437 , TESTIDCODE: 68523LCM60 |
| Open Data Set |
| Plotted Points Statistics |
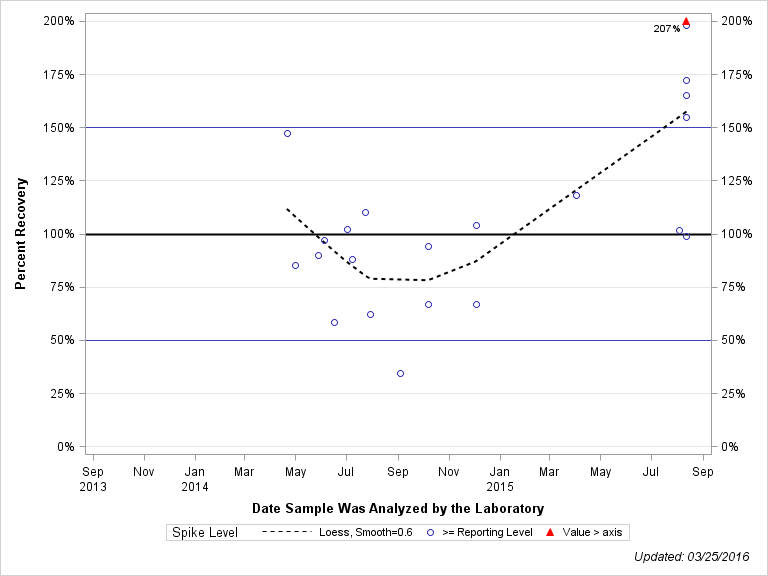
| ACETOCHLOR SULFONIC ACID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 110% | 46% | 100% | 46% |
| Total | 22 | 110% | 46% | 100% | 46% |
| Sample Statistics |
| ACETOCHLOR SULFONIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 81% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 4 | 15% | 4 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| BQS OBSP Determination: |
| ACETOCHLOR SULFYNILACETIC ACID |
| METHOD: 2437 , TESTIDCODE: 68524LCM60 |
| Open Data Set |

| Sample Statistics |
| ACETOCHLOR SULFYNILACETIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| ALACHLOR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 28 | 103% | 23% | 100% | 21% |
| Total | 28 | 103% | 23% | 100% | 21% |
| Sample Statistics |
| ALACHLOR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 28 | 100% | Spiked |
| Estimated Values | 3 | 11% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 0 | 0% | 0 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 0 | 0% | 0 out of 20 |
| Plotted Points Statistics |
| ALACHLOR OA |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 27 | 109% | 22% | 108% | 18% |
| Total | 27 | 109% | 22% | 108% | 18% |
| Sample Statistics |
| ALACHLOR OA |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 27 | 96% | Spiked |
| Estimated Values | 2 | 7% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 0 | 0% | 0 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 4 | 20% | 4 out of 20 |
| Plotted Points Statistics |
| ALACHLOR SULFONIC ACID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 1 | 67% | . | 67% | 0% |
| >= Reporting Level | 20 | 79% | 22% | 77% | 15% |
| Total | 21 | 78% | 21% | 76% | 13% |
| Sample Statistics |
| ALACHLOR SULFONIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 78% | Spiked |
| Estimated Values | 21 | 78% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 4 | 15% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 2 | 7% | 2 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| BQS OBSP Determination: |
| ALACHLOR SULFYNILACETIC ACID |
| METHOD: 2437 , TESTIDCODE: 68527LCM60 |
| Open Data Set |

| Sample Statistics |
| ALACHLOR SULFYNILACETIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| ALDICARB |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 82% | 16% | 82% | 15% |
| Total | 26 | 82% | 16% | 82% | 15% |
| Sample Statistics |
| ALDICARB |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 100% | Spiked |
| Estimated Values | 4 | 15% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 0 | 0% | 0 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| ALDICARB SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 90% | 17% | 87% | 13% |
| Total | 22 | 90% | 17% | 87% | 13% |
| Sample Statistics |
| ALDICARB SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 100% | Spiked |
| Estimated Values | 3 | 14% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 0 | 0% | 0 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| ALDICARB SULFOXIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 24 | 69% | 19% | 67% | 22% |
| Total | 24 | 69% | 19% | 67% | 22% |
| Sample Statistics |
| ALDICARB SULFOXIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 24 | 100% | Spiked |
| Estimated Values | 21 | 88% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 0 | 0% | 0 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 4 | 17% | 4 out of 24 |
| Plotted Points Statistics |
| AMETRYN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 84% | 11% | 83% | 8% |
| Total | 21 | 84% | 11% | 83% | 8% |
| Sample Statistics |
| AMETRYN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 88% | Spiked |
| Estimated Values | 5 | 21% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 8% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 1 | 4% | 1 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 2 | 8% | 2 out of 24 |
| Plotted Points Statistics |
| ASULAM |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 19 | 120% | 35% | 122% | 32% |
| Total | 19 | 120% | 35% | 122% | 32% |
| Sample Statistics |
| ASULAM |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 19 | 86% | Spiked |
| Estimated Values | 19 | 86% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 3 | 14% | 3 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| ATRAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 30 | 98% | 12% | 95% | 12% |
| Total | 30 | 98% | 12% | 95% | 12% |
| Sample Statistics |
| ATRAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 30 | 100% | Spiked |
| Estimated Values | 3 | 10% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 30 | . | |
| False Negatives | 0 | 0% | 0 out of 30 |
| Not Spiked | 18 | . | |
| False Positives | 1 | 6% | 1 out of 18 |
| Plotted Points Statistics |
| AZINPHOS-METHYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 27 | 96% | 28% | 86% | 14% |
| Total | 27 | 96% | 28% | 86% | 14% |
| Sample Statistics |
| AZINPHOS-METHYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 27 | 96% | Spiked |
| Estimated Values | 7 | 25% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 1 | 4% | 1 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 0 | 0% | 0 out of 20 |
| Plotted Points Statistics |
| AZINPHOS-METHYL-OXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 86% | 17% | 82% | 18% |
| Total | 26 | 86% | 17% | 82% | 18% |
| Sample Statistics |
| AZINPHOS-METHYL-OXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 96% | Spiked |
| Estimated Values | 4 | 15% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 1 | 4% | 1 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| Plotted Points Statistics |
| AZOXYSTROBIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 81% | 11% | 80% | 6% |
| Total | 26 | 81% | 11% | 80% | 6% |
| Sample Statistics |
| AZOXYSTROBIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 100% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 0 | 0% | 0 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| BENTAZONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 24 | 105% | 24% | 104% | 21% |
| Total | 24 | 105% | 24% | 104% | 21% |
| Sample Statistics |
| BENTAZONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 24 | 100% | Spiked |
| Estimated Values | 3 | 13% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 0 | 0% | 0 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 0 | 0% | 0 out of 24 |
| Plotted Points Statistics |
| BIFENTHRIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 19 | 32% | 14% | 31% | 11% |
| Total | 19 | 32% | 14% | 31% | 11% |
| Sample Statistics |
| BIFENTHRIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 19 | 95% | Spiked |
| Estimated Values | 19 | 95% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 20 | . | |
| False Negatives | 1 | 5% | 1 out of 20 |
| Not Spiked | 28 | . | |
| False Positives | 0 | 0% | 0 out of 28 |
| Plotted Points Statistics |
| BROMACIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 92% | 21% | 92% | 16% |
| Total | 25 | 92% | 21% | 92% | 16% |
| Sample Statistics |
| BROMACIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 96% | Spiked |
| Estimated Values | 4 | 15% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 1 | 4% | 1 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| BROMOXYNIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 106% | 18% | 110% | 15% |
| Total | 22 | 106% | 18% | 110% | 15% |
| Sample Statistics |
| BROMOXYNIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 100% | Spiked |
| Estimated Values | 2 | 9% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 0 | 0% | 0 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| BUTRALIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 16 | 81% | 11% | 82% | 10% |
| Total | 16 | 81% | 11% | 82% | 10% |
| Sample Statistics |
| BUTRALIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 16 | 89% | Spiked |
| Estimated Values | 16 | 89% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 6% | Spiked |
| Spiked | 18 | . | |
| False Negatives | 1 | 6% | 1 out of 18 |
| Not Spiked | 30 | . | |
| False Positives | 0 | 0% | 0 out of 30 |
| Plotted Points Statistics |
| BUTYLATE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 12 | 78% | 17% | 84% | 13% |
| Total | 12 | 78% | 17% | 84% | 13% |
| Sample Statistics |
| BUTYLATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 12 | 48% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 10 | 40% | Spiked |
| Spiked | 25 | . | |
| False Negatives | 3 | 12% | 3 out of 25 |
| Not Spiked | 23 | . | |
| False Positives | 0 | 0% | 0 out of 23 |
| Plotted Points Statistics |
| CARBARYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 33 | 107% | 22% | 106% | 15% |
| Total | 33 | 107% | 22% | 106% | 15% |
| Sample Statistics |
| CARBARYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 33 | 97% | Spiked |
| Estimated Values | 3 | 9% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 34 | . | |
| False Negatives | 1 | 3% | 1 out of 34 |
| Not Spiked | 14 | . | |
| False Positives | 0 | 0% | 0 out of 14 |
| Plotted Points Statistics |
| CARBENDAZIM |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 237% | 26% | 239% | 26% |
| Total | 22 | 237% | 26% | 239% | 26% |
| Sample Statistics |
| CARBENDAZIM |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 92% | Spiked |
| Estimated Values | 22 | 92% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 8% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 0 | 0% | 0 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 4 | 17% | 4 out of 24 |
| Plotted Points Statistics |
| CARBOFURAN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 30 | 218% | 364% | 101% | 14% |
| Total | 30 | 218% | 364% | 101% | 14% |
| Sample Statistics |
| CARBOFURAN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 30 | 100% | Spiked |
| Estimated Values | 2 | 7% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 30 | . | |
| False Negatives | 0 | 0% | 0 out of 30 |
| Not Spiked | 18 | . | |
| False Positives | 4 | 22% | 4 out of 18 |
| Sample Statistics |
| CARBOXY MOLINATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| CHLORIMURON-ETHYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 15 | 57% | 27% | 57% | 35% |
| Total | 15 | 57% | 27% | 57% | 35% |
| Sample Statistics |
| CHLORIMURON-ETHYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 15 | 60% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 25 | . | |
| False Negatives | 10 | 40% | 10 out of 25 |
| Not Spiked | 23 | . | |
| False Positives | 0 | 0% | 0 out of 23 |
| Sample Statistics |
| CHLOROSULFONAMIDE ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| CHLORPYRIFOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 29 | 86% | 19% | 82% | 18% |
| Total | 29 | 86% | 19% | 82% | 18% |
| Sample Statistics |
| CHLORPYRIFOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 29 | 100% | Spiked |
| Estimated Values | 5 | 17% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 29 | . | |
| False Negatives | 0 | 0% | 0 out of 29 |
| Not Spiked | 19 | . | |
| False Positives | 0 | 0% | 0 out of 19 |
| Plotted Points Statistics |
| CHLORPYRIFOS OXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 18 | 107% | 41% | 114% | 43% |
| Total | 18 | 107% | 41% | 114% | 43% |
| Sample Statistics |
| CHLORPYRIFOS OXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 18 | 75% | Spiked |
| Estimated Values | 15 | 63% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 6 | 25% | 6 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 0 | 0% | 0 out of 24 |
| Plotted Points Statistics |
| CHLORSULFURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 16 | 94% | 25% | 100% | 17% |
| Total | 16 | 94% | 25% | 100% | 17% |
| Sample Statistics |
| CHLORSULFURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 16 | 100% | Spiked |
| Estimated Values | 1 | 6% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 16 | . | |
| False Negatives | 0 | 0% | 0 out of 16 |
| Not Spiked | 32 | . | |
| False Positives | 4 | 13% | 4 out of 32 |
| BQS OBSP Determination: |
| CIS-BIFENTHRIN ACID/CIS-CYHALOTHRIN ACID/CIS-TEFLUTHRIN ACID |
| METHOD: 2437 , TESTIDCODE: 68553LCM60 |
| Open Data Set |

| Sample Statistics |
| CIS-BIFENTHRIN ACID/CIS-CYHALOTHRIN ACID/CIS-TEFLUTHRIN ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| CIS-PERMETHRIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 59% | 15% | 59% | 12% |
| Total | 21 | 59% | 15% | 59% | 12% |
| Sample Statistics |
| CIS-PERMETHRIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 72% | Spiked |
| Estimated Values | 15 | 52% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 3% | Spiked |
| Spiked | 29 | . | |
| False Negatives | 7 | 24% | 7 out of 29 |
| Not Spiked | 19 | . | |
| False Positives | 0 | 0% | 0 out of 19 |
| Plotted Points Statistics |
| CYANAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 19 | 99% | 52% | 92% | 40% |
| Total | 19 | 99% | 52% | 92% | 40% |
| Sample Statistics |
| CYANAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 19 | 63% | Spiked |
| Estimated Values | 9 | 30% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 6 | 20% | Spiked |
| Spiked | 30 | . | |
| False Negatives | 5 | 17% | 5 out of 30 |
| Not Spiked | 18 | . | |
| False Positives | 0 | 0% | 0 out of 18 |
| Plotted Points Statistics |
| DACTHAL MONOACID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 2 | 78% | 26% | 78% | 27% |
| >= Reporting Level | 15 | 105% | 58% | 92% | 19% |
| Total | 17 | 102% | 55% | 92% | 19% |
| Sample Statistics |
| DACTHAL MONOACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 17 | 85% | Spiked |
| Estimated Values | 17 | 85% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 10% | Spiked |
| Spiked | 20 | . | |
| False Negatives | 1 | 5% | 1 out of 20 |
| Not Spiked | 28 | . | |
| False Positives | 4 | 14% | 4 out of 28 |
| Sample Statistics |
| DECHLOROFIPRONIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| DECHLOROMETOLACHLOR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 6 | 86% | 5% | 86% | 5% |
| Total | 6 | 86% | 5% | 86% | 5% |
| Sample Statistics |
| DECHLOROMETOLACHLOR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 6 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 6 | . | |
| False Negatives | 0 | 0% | 0 out of 6 |
| Not Spiked | 42 | . | |
| False Positives | 0 | 0% | 0 out of 42 |
| Plotted Points Statistics |
| DEETHYLATRAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 32 | 106% | 18% | 103% | 19% |
| Total | 32 | 106% | 18% | 103% | 19% |
| Sample Statistics |
| DEETHYLATRAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 32 | 100% | Spiked |
| Estimated Values | 7 | 22% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 32 | . | |
| False Negatives | 0 | 0% | 0 out of 32 |
| Not Spiked | 16 | . | |
| False Positives | 0 | 0% | 0 out of 16 |
| BQS OBSP Determination: |
| DEETHYLDEISOPROPYLATRAZINE |
| METHOD: 2437 , TESTIDCODE: 68547LCM60 |
| Open Data Set |
| Plotted Points Statistics |
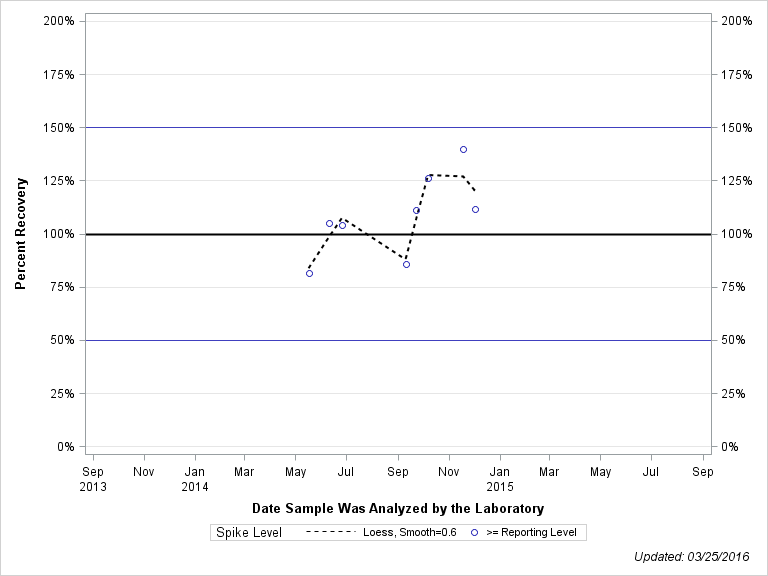
| DEETHYLDEISOPROPYLATRAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 8 | 108% | 19% | 108% | 18% |
| Total | 8 | 108% | 19% | 108% | 18% |
| Sample Statistics |
| DEETHYLDEISOPROPYLATRAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 8 | 36% | Spiked |
| Estimated Values | 2 | 9% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 5% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 13 | 59% | 13 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 4 | 15% | 4 out of 26 |
| Sample Statistics |
| DEIODO FLUBENDIAMIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| DEISOPROPYLATRAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 88% | 17% | 89% | 19% |
| Total | 22 | 88% | 17% | 89% | 19% |
| Sample Statistics |
| DEISOPROPYLATRAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 92% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 2 | 8% | 2 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 0 | 0% | 0 out of 24 |
| Sample Statistics |
| DEISOPROPYL PROMETRYN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| DEMETHYL FLUOMETURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 11 | 101% | 16% | 104% | 23% |
| Total | 11 | 101% | 16% | 104% | 23% |
| Sample Statistics |
| DEMETHYL FLUOMETURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 11 | 85% | Spiked |
| Estimated Values | 1 | 8% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 13 | . | |
| False Negatives | 2 | 15% | 2 out of 13 |
| Not Spiked | 35 | . | |
| False Positives | 0 | 0% | 0 out of 35 |
| Sample Statistics |
| DEMETHYL HEXAZINONE B |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| DEMETHYL NORFLURAZON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 10 | 115% | 53% | 90% | 57% |
| Total | 10 | 115% | 53% | 90% | 57% |
| Sample Statistics |
| DEMETHYL NORFLURAZON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 10 | 83% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 17% | Spiked |
| Spiked | 12 | . | |
| False Negatives | 0 | 0% | 0 out of 12 |
| Not Spiked | 36 | . | |
| False Positives | 0 | 0% | 0 out of 36 |
| BQS OBSP Determination: |
| DESAMINO-DIKETO METRIBUZIN |
| METHOD: 2437 , TESTIDCODE: 68569LCM60 |
| Open Data Set |
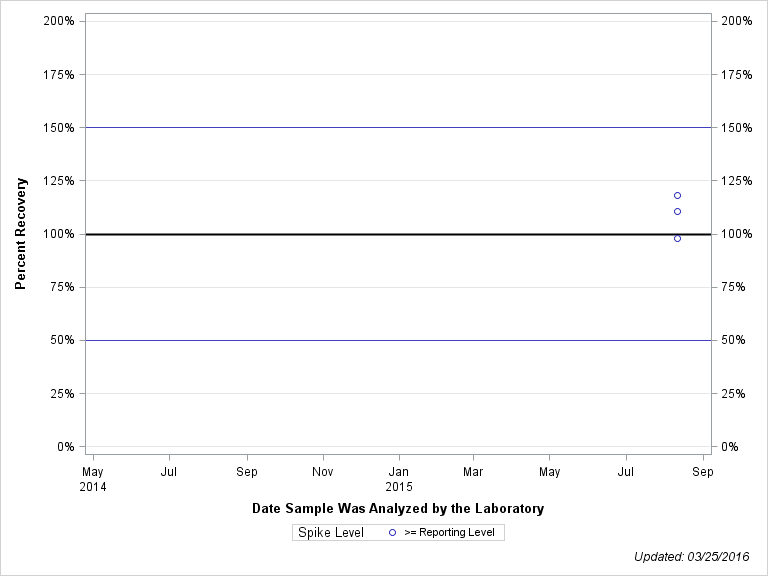
| Plotted Points Statistics |
| DESAMINO-DIKETO METRIBUZIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 3 | 109% | 10% | 110% | 15% |
| Total | 3 | 109% | 10% | 110% | 15% |
| Sample Statistics |
| DESAMINO-DIKETO METRIBUZIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 3 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 3 | . | |
| False Negatives | 0 | 0% | 0 out of 3 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| Plotted Points Statistics |
| DESULFINYLFIPRONIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 92% | 15% | 91% | 17% |
| Total | 21 | 92% | 15% | 91% | 17% |
| Sample Statistics |
| DESULFINYLFIPRONIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 95% | Spiked |
| Estimated Values | 1 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 1 | 5% | 1 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| BQS OBSP Determination: |
| DESULFINYLFIPRONIL AMIDE |
| METHOD: 2437 , TESTIDCODE: 68570LCM60 |
| Open Data Set |

| Sample Statistics |
| DESULFINYLFIPRONIL AMIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| DIAZINON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 24 | 106% | 21% | 111% | 17% |
| Total | 24 | 106% | 21% | 111% | 17% |
| Sample Statistics |
| DIAZINON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 24 | 92% | Spiked |
| Estimated Values | 19 | 73% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 1 | 4% | 1 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| DIAZINON OXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 73% | 15% | 69% | 18% |
| Total | 25 | 73% | 15% | 69% | 18% |
| Sample Statistics |
| DIAZINON OXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 93% | Spiked |
| Estimated Values | 25 | 93% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 2 | 7% | 2 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 1 | 5% | 1 out of 21 |
| Plotted Points Statistics |
| DICAMBA |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 17 | 93% | 34% | 86% | 20% |
| Total | 17 | 93% | 34% | 86% | 20% |
| Sample Statistics |
| DICAMBA |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 17 | 77% | Spiked |
| Estimated Values | 17 | 77% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 5 | 23% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 0 | 0% | 0 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| DICHLORVOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 127% | 39% | 118% | 16% |
| Total | 26 | 127% | 39% | 118% | 16% |
| Sample Statistics |
| DICHLORVOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 96% | Spiked |
| Estimated Values | 22 | 81% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 0 | 0% | 0 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| Plotted Points Statistics |
| DICROTOPHOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 116% | 27% | 109% | 19% |
| Total | 26 | 116% | 27% | 109% | 19% |
| Sample Statistics |
| DICROTOPHOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 96% | Spiked |
| Estimated Values | 13 | 48% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 0 | 0% | 0 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| Sample Statistics |
| DIDEMETHYL HEXAZINONE F |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| DIFLUBENZURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 98% | 14% | 99% | 9% |
| Total | 22 | 98% | 14% | 99% | 9% |
| Sample Statistics |
| DIFLUBENZURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 100% | Spiked |
| Estimated Values | 4 | 18% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 0 | 0% | 0 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 1 | 4% | 1 out of 26 |
| Plotted Points Statistics |
| DIFLUFENZOPYR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 15 | 91% | 34% | 88% | 35% |
| Total | 15 | 91% | 34% | 88% | 35% |
| Sample Statistics |
| DIFLUFENZOPYR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 15 | 94% | Spiked |
| Estimated Values | 1 | 6% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 16 | . | |
| False Negatives | 1 | 6% | 1 out of 16 |
| Not Spiked | 32 | . | |
| False Positives | 4 | 13% | 4 out of 32 |
| BQS OBSP Determination: |
| DIKETONITRILE-ISOXAFLUTOLE |
| METHOD: 2437 , TESTIDCODE: 68578LCM60 |
| Open Data Set |
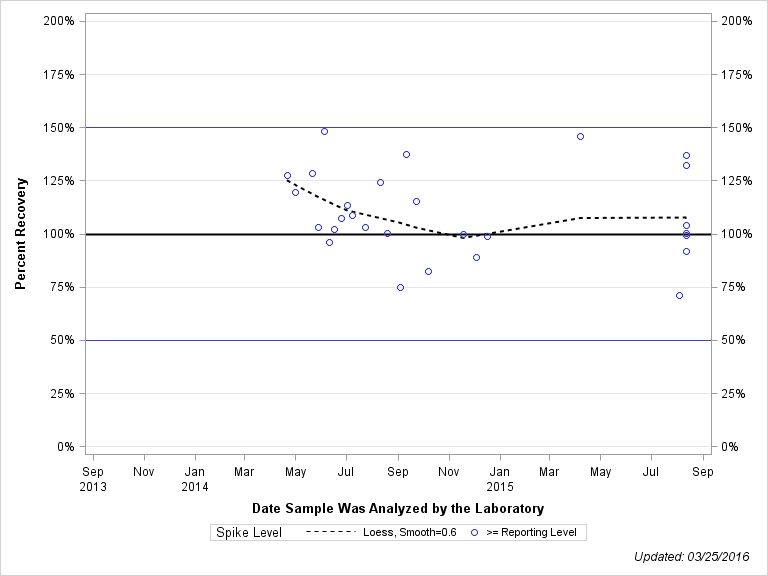
| Plotted Points Statistics |
| DIKETONITRILE-ISOXAFLUTOLE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 28 | 109% | 20% | 104% | 20% |
| Total | 28 | 109% | 20% | 104% | 20% |
| Sample Statistics |
| DIKETONITRILE-ISOXAFLUTOLE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 28 | 100% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 0 | 0% | 0 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 3 | 15% | 3 out of 20 |
| Plotted Points Statistics |
| DIMETHENAMID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 28 | 91% | 20% | 93% | 14% |
| Total | 28 | 91% | 20% | 93% | 14% |
| Sample Statistics |
| DIMETHENAMID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 28 | 97% | Spiked |
| Estimated Values | 2 | 7% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 29 | . | |
| False Negatives | 1 | 3% | 1 out of 29 |
| Not Spiked | 19 | . | |
| False Positives | 1 | 5% | 1 out of 19 |
| BQS OBSP Determination: |
| DIMETHENAMID OXANILIC ACID |
| METHOD: 2437 , TESTIDCODE: 68581LCM60 |
| Open Data Set |
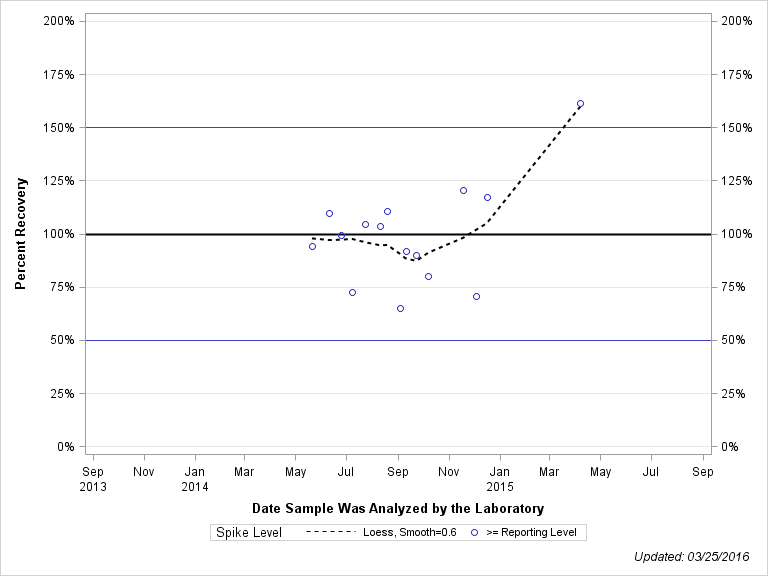
| Plotted Points Statistics |
| DIMETHENAMID OXANILIC ACID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 15 | 99% | 24% | 100% | 23% |
| Total | 15 | 99% | 24% | 100% | 23% |
| Sample Statistics |
| DIMETHENAMID OXANILIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 15 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 15 | . | |
| False Negatives | 0 | 0% | 0 out of 15 |
| Not Spiked | 33 | . | |
| False Positives | 1 | 3% | 1 out of 33 |
| Sample Statistics |
| DIMETHENAMID SAA |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| BQS OBSP Determination: |
| DIMETHENAMID SULFONIC ACID |
| METHOD: 2437 , TESTIDCODE: 68582LCM60 |
| Open Data Set |
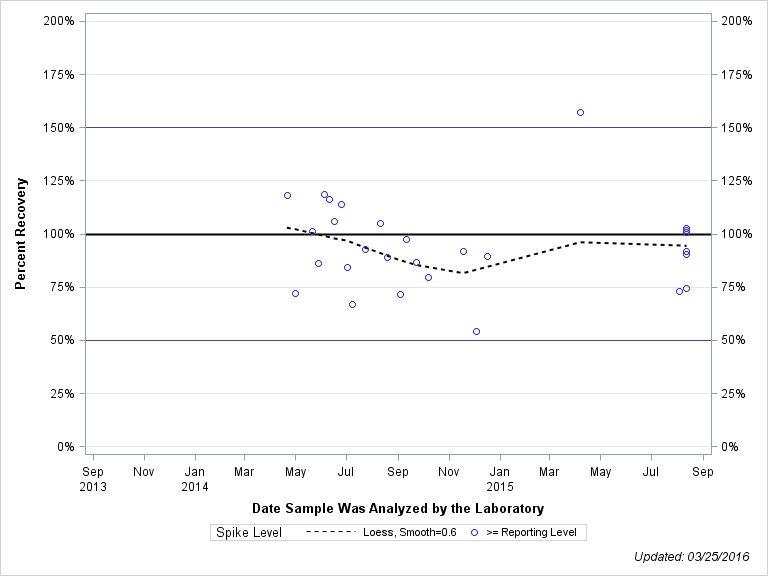
| Plotted Points Statistics |
| DIMETHENAMID SULFONIC ACID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 28 | 94% | 20% | 92% | 16% |
| Total | 28 | 94% | 20% | 92% | 16% |
| Sample Statistics |
| DIMETHENAMID SULFONIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 28 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 0 | 0% | 0 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 0 | 0% | 0 out of 20 |
| Plotted Points Statistics |
| DIMETHOATE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 115% | 33% | 109% | 24% |
| Total | 25 | 115% | 33% | 109% | 24% |
| Sample Statistics |
| DIMETHOATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 100% | Spiked |
| Estimated Values | 6 | 24% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 25 | . | |
| False Negatives | 0 | 0% | 0 out of 25 |
| Not Spiked | 23 | . | |
| False Positives | 0 | 0% | 0 out of 23 |
| BQS OBSP Determination: |
| DIMETHOATE, OXYGEN ANALOG |
| METHOD: 2437 , TESTIDCODE: 68661LCM60 |
| Open Data Set |
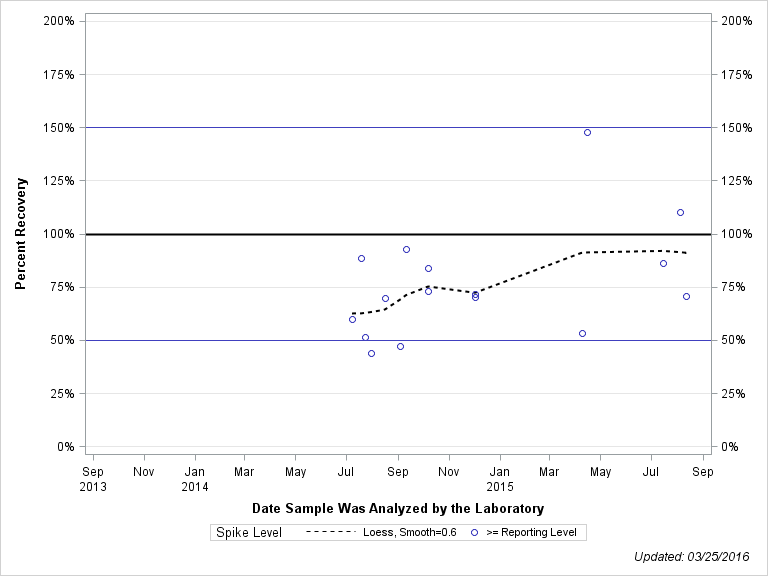
| Plotted Points Statistics |
| DIMETHOATE, OXYGEN ANALOG |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 16 | 76% | 26% | 71% | 23% |
| Total | 16 | 76% | 26% | 71% | 23% |
| Sample Statistics |
| DIMETHOATE, OXYGEN ANALOG |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 16 | 84% | Spiked |
| Estimated Values | 4 | 21% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 19 | . | |
| False Negatives | 3 | 16% | 3 out of 19 |
| Not Spiked | 29 | . | |
| False Positives | 0 | 0% | 0 out of 29 |
| Plotted Points Statistics |
| DISULFOTON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 63% | 28% | 69% | 23% |
| Total | 25 | 63% | 28% | 69% | 23% |
| Sample Statistics |
| DISULFOTON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 89% | Spiked |
| Estimated Values | 7 | 25% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 3 | 11% | 3 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 0 | 0% | 0 out of 20 |
| Plotted Points Statistics |
| DISULFOTON OXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 17 | 69% | 22% | 64% | 21% |
| Total | 17 | 69% | 22% | 64% | 21% |
| Sample Statistics |
| DISULFOTON OXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 17 | 94% | Spiked |
| Estimated Values | 4 | 22% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 18 | . | |
| False Negatives | 1 | 6% | 1 out of 18 |
| Not Spiked | 30 | . | |
| False Positives | 0 | 0% | 0 out of 30 |
| Plotted Points Statistics |
| DISULFOTON OXON SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 1 | 71% | . | 71% | 0% |
| Total | 1 | 71% | . | 71% | 0% |
| Sample Statistics |
| DISULFOTON OXON SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 1 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 1 | . | |
| False Negatives | 0 | 0% | 0 out of 1 |
| Not Spiked | 47 | . | |
| False Positives | 0 | 0% | 0 out of 47 |
| BQS OBSP Determination: |
| DISULFOTON OXON SULFOXIDE |
| METHOD: 2437 , TESTIDCODE: 68587LCM60 |
| Open Data Set |
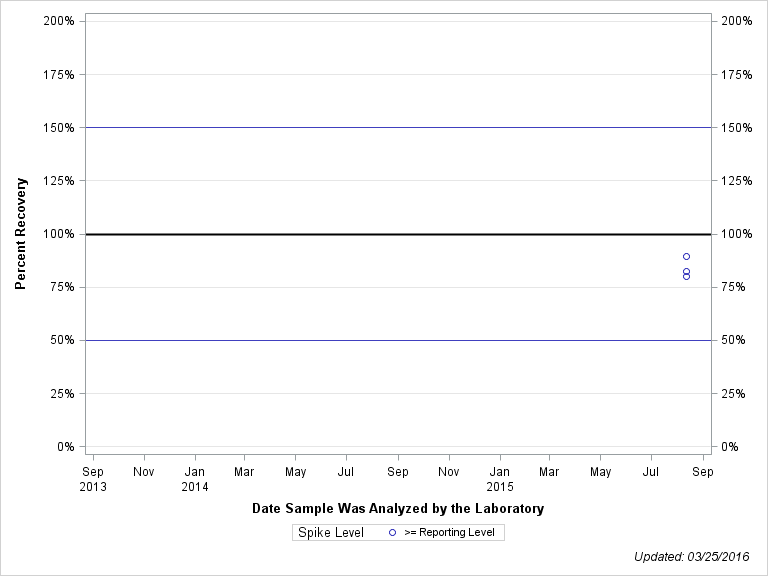
| Plotted Points Statistics |
| DISULFOTON OXON SULFOXIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 3 | 84% | 5% | 83% | 7% |
| Total | 3 | 84% | 5% | 83% | 7% |
| Sample Statistics |
| DISULFOTON OXON SULFOXIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 3 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 3 | . | |
| False Negatives | 0 | 0% | 0 out of 3 |
| Not Spiked | 45 | . | |
| False Positives | 8 | 18% | 8 out of 45 |
| Plotted Points Statistics |
| DISULFOTON SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 20 | 84% | 17% | 82% | 14% |
| Total | 20 | 84% | 17% | 82% | 14% |
| Sample Statistics |
| DISULFOTON SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 20 | 100% | Spiked |
| Estimated Values | 5 | 25% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 20 | . | |
| False Negatives | 0 | 0% | 0 out of 20 |
| Not Spiked | 28 | . | |
| False Positives | 2 | 7% | 2 out of 28 |
| Plotted Points Statistics |
| DISULFOTON SULFOXIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 18 | 175% | 118% | 156% | 82% |
| Total | 18 | 175% | 118% | 156% | 82% |
| Plotted Points Statistics |
| DIURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 92% | 16% | 94% | 9% |
| Total | 21 | 92% | 16% | 94% | 9% |
| Sample Statistics |
| DIURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 81% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 8% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 3 | 12% | 3 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| EPTC |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 4 | 77% | 39% | 64% | 34% |
| >= Reporting Level | 21 | 91% | 27% | 93% | 19% |
| Total | 25 | 89% | 29% | 91% | 22% |
| Sample Statistics |
| EPTC |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 68% | Spiked |
| Estimated Values | 1 | 3% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 11 | 30% | Spiked |
| Spiked | 37 | . | |
| False Negatives | 1 | 3% | 1 out of 37 |
| Not Spiked | 11 | . | |
| False Positives | 0 | 0% | 0 out of 11 |
| Sample Statistics |
| EPTC DEGRADATE R248722 |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| ETHOPROPHOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 30 | 101% | 18% | 97% | 21% |
| Total | 30 | 101% | 18% | 97% | 21% |
| Sample Statistics |
| ETHOPROPHOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 30 | 100% | Spiked |
| Estimated Values | 4 | 13% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 30 | . | |
| False Negatives | 0 | 0% | 0 out of 30 |
| Not Spiked | 18 | . | |
| False Positives | 0 | 0% | 0 out of 18 |
| Plotted Points Statistics |
| ETOXAZOLE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 88% | 12% | 84% | 13% |
| Total | 21 | 88% | 12% | 84% | 13% |
| Sample Statistics |
| ETOXAZOLE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 95% | Spiked |
| Estimated Values | 5 | 23% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 1 | 5% | 1 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| FAMOXADONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 10 | 35% | 20% | 41% | 24% |
| Total | 10 | 35% | 20% | 41% | 24% |
| Sample Statistics |
| FAMOXADONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 10 | 59% | Spiked |
| Estimated Values | 10 | 59% | Spiked |
| Deleted Values | 4 | 8% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 18 | . | |
| False Negatives | 7 | 41% | 7 out of 17 |
| Not Spiked | 30 | . | |
| False Positives | 0 | 0% | 0 out of 27 |
| Plotted Points Statistics |
| FENAMIPHOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 61% | 16% | 60% | 14% |
| Total | 22 | 61% | 16% | 60% | 14% |
| Sample Statistics |
| FENAMIPHOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 92% | Spiked |
| Estimated Values | 18 | 75% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 2 | 8% | 2 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 0 | 0% | 0 out of 24 |
| Plotted Points Statistics |
| FENAMIPHOS SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 86% | 19% | 84% | 20% |
| Total | 26 | 86% | 19% | 84% | 20% |
| Sample Statistics |
| FENAMIPHOS SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 96% | Spiked |
| Estimated Values | 6 | 22% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 0 | 0% | 0 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| Plotted Points Statistics |
| FENAMIPHOS SULFOXIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 24 | 109% | 31% | 103% | 25% |
| Total | 24 | 109% | 31% | 103% | 25% |
| Sample Statistics |
| FENAMIPHOS SULFOXIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 24 | 96% | Spiked |
| Estimated Values | 5 | 20% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 25 | . | |
| False Negatives | 1 | 4% | 1 out of 25 |
| Not Spiked | 23 | . | |
| False Positives | 3 | 13% | 3 out of 23 |
| Plotted Points Statistics |
| FENBUTATIN OXIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 3 | 38% | 2% | 39% | 3% |
| Total | 3 | 38% | 2% | 39% | 3% |
| Sample Statistics |
| FENBUTATIN OXIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 3 | 17% | Spiked |
| Estimated Values | 3 | 17% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 4 | 22% | Spiked |
| Spiked | 18 | . | |
| False Negatives | 11 | 61% | 11 out of 18 |
| Not Spiked | 30 | . | |
| False Positives | 0 | 0% | 0 out of 30 |
| Plotted Points Statistics |
| FENTIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 3 | 103% | 2% | 102% | 3% |
| Total | 3 | 103% | 2% | 102% | 3% |
| Sample Statistics |
| FENTIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 3 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 3 | . | |
| False Negatives | 0 | 0% | 0 out of 3 |
| Not Spiked | 45 | . | |
| False Positives | 0 | 0% | 0 out of 45 |
| Plotted Points Statistics |
| FIPRONIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 30 | 92% | 17% | 91% | 18% |
| Total | 30 | 92% | 17% | 91% | 18% |
| Sample Statistics |
| FIPRONIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 30 | 100% | Spiked |
| Estimated Values | 2 | 7% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 30 | . | |
| False Negatives | 0 | 0% | 0 out of 30 |
| Not Spiked | 18 | . | |
| False Positives | 0 | 0% | 0 out of 18 |
| Sample Statistics |
| FIPRONIL AMIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| FIPRONIL SULFIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 28 | 108% | 27% | 104% | 17% |
| Total | 28 | 108% | 27% | 104% | 17% |
| Sample Statistics |
| FIPRONIL SULFIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 28 | 100% | Spiked |
| Estimated Values | 6 | 21% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 0 | 0% | 0 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 0 | 0% | 0 out of 20 |
| Sample Statistics |
| FIPRONIL SULFONATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| FIPRONIL SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 96% | 23% | 94% | 13% |
| Total | 26 | 96% | 23% | 94% | 13% |
| Sample Statistics |
| FIPRONIL SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 100% | Spiked |
| Estimated Values | 5 | 19% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 0 | 0% | 0 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| FLUBENDIAMIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 27 | 97% | 26% | 89% | 25% |
| Total | 27 | 97% | 26% | 89% | 25% |
| Sample Statistics |
| FLUBENDIAMIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 27 | 96% | Spiked |
| Estimated Values | 3 | 11% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 0 | 0% | 0 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 0 | 0% | 0 out of 20 |
| Plotted Points Statistics |
| FLUMETSULAM |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 128% | 21% | 127% | 13% |
| Total | 21 | 128% | 21% | 127% | 13% |
| Sample Statistics |
| FLUMETSULAM |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 95% | Spiked |
| Estimated Values | 21 | 95% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 5% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 0 | 0% | 0 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| FLUOMETURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 18 | 107% | 17% | 104% | 17% |
| Total | 18 | 107% | 17% | 104% | 17% |
| Sample Statistics |
| FLUOMETURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 18 | 82% | Spiked |
| Estimated Values | 2 | 9% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 3 | 14% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 1 | 5% | 1 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| FONOFOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 102% | 28% | 104% | 16% |
| Total | 25 | 102% | 28% | 104% | 16% |
| Sample Statistics |
| FONOFOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 89% | Spiked |
| Estimated Values | 2 | 7% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 2 | 7% | 2 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 0 | 0% | 0 out of 20 |
| Plotted Points Statistics |
| HALOSULFURON-METHYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 18 | 94% | 20% | 100% | 25% |
| Total | 18 | 94% | 20% | 100% | 25% |
| Sample Statistics |
| HALOSULFURON-METHYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 18 | 100% | Spiked |
| Estimated Values | 1 | 6% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 18 | . | |
| False Negatives | 0 | 0% | 0 out of 18 |
| Not Spiked | 30 | . | |
| False Positives | 0 | 0% | 0 out of 30 |
| Plotted Points Statistics |
| HEXAZINONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 91% | 11% | 89% | 10% |
| Total | 25 | 91% | 11% | 89% | 10% |
| Sample Statistics |
| HEXAZINONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 100% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 25 | . | |
| False Negatives | 0 | 0% | 0 out of 25 |
| Not Spiked | 23 | . | |
| False Positives | 0 | 0% | 0 out of 23 |
| BQS OBSP Determination: |
| HEXAZINONE TRANSFORMATION PRODUCT C |
| METHOD: 2437 , TESTIDCODE: 68612LCM60 |
| Open Data Set |

| Sample Statistics |
| HEXAZINONE TRANSFORMATION PRODUCT C |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| BQS OBSP Determination: |
| HEXAZINONE TRANSFORMATION PRODUCT D |
| METHOD: 2437 , TESTIDCODE: 68613LCM60 |
| Open Data Set |

| Sample Statistics |
| HEXAZINONE TRANSFORMATION PRODUCT D |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| BQS OBSP Determination: |
| HEXAZINONE TRANSFORMATION PRODUCT E |
| METHOD: 2437 , TESTIDCODE: 68614LCM60 |
| Open Data Set |

| Sample Statistics |
| HEXAZINONE TRANSFORMATION PRODUCT E |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| BQS OBSP Determination: |
| HEXAZINONE TRANSFORMATION PRODUCT G |
| METHOD: 2437 , TESTIDCODE: 68713LCM60 |
| Open Data Set |

| Sample Statistics |
| HEXAZINONE TRANSFORMATION PRODUCT G |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Sample Statistics |
| HYDROXYACETOCHLOR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 1 | 2% | 1 out of 48 |
| Plotted Points Statistics |
| HYDROXYALACHLOR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 6 | 96% | 10% | 94% | 8% |
| Total | 6 | 96% | 10% | 94% | 8% |
| Sample Statistics |
| HYDROXYALACHLOR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 6 | 100% | Spiked |
| Estimated Values | 1 | 17% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 6 | . | |
| False Negatives | 0 | 0% | 0 out of 6 |
| Not Spiked | 42 | . | |
| False Positives | 0 | 0% | 0 out of 42 |
| Sample Statistics |
| HYDROXYDIAZINON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Sample Statistics |
| HYDROXYFLUOMETURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| HYDROXYMETOLACHLOR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 6 | 86% | 6% | 85% | 8% |
| Total | 6 | 86% | 6% | 85% | 8% |
| Sample Statistics |
| HYDROXYMETOLACHLOR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 6 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 6 | . | |
| False Negatives | 0 | 0% | 0 out of 6 |
| Not Spiked | 42 | . | |
| False Positives | 1 | 2% | 1 out of 42 |
| Sample Statistics |
| HYDROXYPHTHALAZINONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| HYDROXYSIMAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 13 | 86% | 18% | 92% | 7% |
| Total | 13 | 86% | 18% | 92% | 7% |
| Sample Statistics |
| HYDROXYSIMAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 13 | 50% | Spiked |
| Estimated Values | 2 | 8% | Spiked |
| Deleted Values | 2 | 4% | Spiked + Not Spiked |
| Spiked, Censored | 8 | 31% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 5 | 19% | 5 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 1 | 5% | 1 out of 20 |
| Sample Statistics |
| HYDROXYTEBUTHURION |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| BQS OBSP Determination: |
| HYDROXY DIDEMETHYL FLUOMETURON |
| METHOD: 2437 , TESTIDCODE: 68619LCM60 |
| Open Data Set |

| Sample Statistics |
| HYDROXY DIDEMETHYL FLUOMETURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| BQS OBSP Determination: |
| HYDROXY MONODEMETHYL FLUOMETURON |
| METHOD: 2437 , TESTIDCODE: 68617LCM60 |
| Open Data Set |

| Sample Statistics |
| HYDROXY MONODEMETHYL FLUOMETURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| IMAZAMOX |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 15 | 133% | 24% | 136% | 20% |
| Total | 15 | 133% | 24% | 136% | 20% |
| Sample Statistics |
| IMAZAMOX |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 15 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 15 | . | |
| False Negatives | 0 | 0% | 0 out of 15 |
| Not Spiked | 33 | . | |
| False Positives | 3 | 9% | 3 out of 33 |
| Plotted Points Statistics |
| IMAZAQUIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 112% | 17% | 115% | 17% |
| Total | 26 | 112% | 17% | 115% | 17% |
| Sample Statistics |
| IMAZAQUIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 100% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 0 | 0% | 0 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| IMAZETHAPYR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 121% | 22% | 126% | 23% |
| Total | 22 | 121% | 22% | 126% | 23% |
| Sample Statistics |
| IMAZETHAPYR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 100% | Spiked |
| Estimated Values | 14 | 64% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 0 | 0% | 0 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| IMIDACLOPRID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 103% | 16% | 101% | 14% |
| Total | 25 | 103% | 16% | 101% | 14% |
| Sample Statistics |
| IMIDACLOPRID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 96% | Spiked |
| Estimated Values | 5 | 19% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 1 | 4% | 1 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| INDOXACARB |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 16 | 110% | 29% | 104% | 35% |
| Total | 16 | 110% | 29% | 104% | 35% |
| Sample Statistics |
| INDOXACARB |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 16 | 94% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 17 | . | |
| False Negatives | 1 | 6% | 1 out of 17 |
| Not Spiked | 31 | . | |
| False Positives | 0 | 0% | 0 out of 31 |
| Plotted Points Statistics |
| ISOXAFLUTOLE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 16 | 88% | 32% | 83% | 38% |
| Total | 16 | 88% | 32% | 83% | 38% |
| Sample Statistics |
| ISOXAFLUTOLE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 16 | 94% | Spiked |
| Estimated Values | 1 | 6% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 6% | Spiked |
| Spiked | 17 | . | |
| False Negatives | 0 | 0% | 0 out of 17 |
| Not Spiked | 31 | . | |
| False Positives | 0 | 0% | 0 out of 31 |
| BQS OBSP Determination: |
| ISOXAFLUTOLE ACID METABOLITE RPA 203328 |
| METHOD: 2437 , TESTIDCODE: 68633LCM60 |
| Open Data Set |
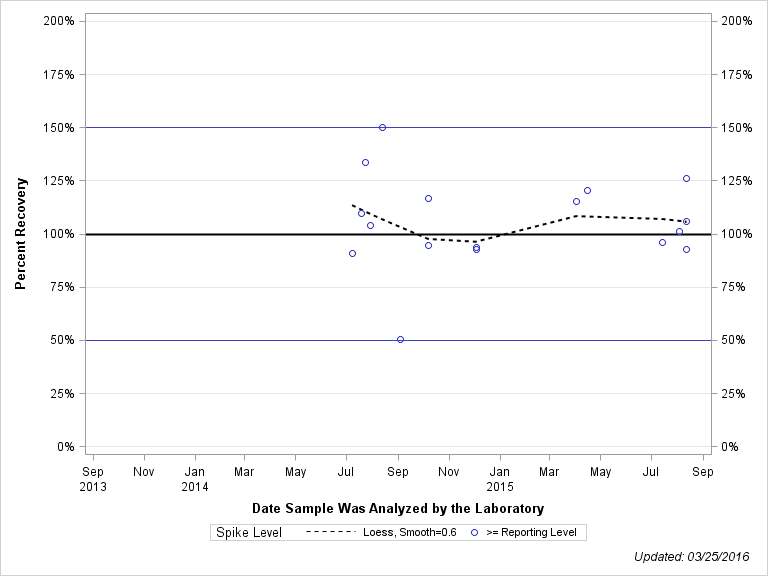
| Plotted Points Statistics |
| ISOXAFLUTOLE ACID METABOLITE RPA 203328 |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 17 | 106% | 22% | 104% | 17% |
| Total | 17 | 106% | 22% | 104% | 17% |
| Sample Statistics |
| ISOXAFLUTOLE ACID METABOLITE RPA 203328 |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 17 | 89% | Spiked |
| Estimated Values | 1 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 11% | Spiked |
| Spiked | 19 | . | |
| False Negatives | 0 | 0% | 0 out of 19 |
| Not Spiked | 29 | . | |
| False Positives | 0 | 0% | 0 out of 29 |
| Plotted Points Statistics |
| KRESOXIM-METHYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 17 | 79% | 11% | 81% | 11% |
| Total | 17 | 79% | 11% | 81% | 11% |
| Sample Statistics |
| KRESOXIM-METHYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 17 | 85% | Spiked |
| Estimated Values | 3 | 15% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 20 | . | |
| False Negatives | 3 | 15% | 3 out of 20 |
| Not Spiked | 28 | . | |
| False Positives | 0 | 0% | 0 out of 28 |
| Plotted Points Statistics |
| LACTOFEN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 15 | 122% | 43% | 118% | 62% |
| Total | 15 | 122% | 43% | 118% | 62% |
| Sample Statistics |
| LACTOFEN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 15 | 79% | Spiked |
| Estimated Values | 15 | 79% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 11% | Spiked |
| Spiked | 19 | . | |
| False Negatives | 2 | 11% | 2 out of 19 |
| Not Spiked | 29 | . | |
| False Positives | 0 | 0% | 0 out of 29 |
| Plotted Points Statistics |
| LINURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 28 | 99% | 21% | 98% | 16% |
| Total | 28 | 99% | 21% | 98% | 16% |
| Sample Statistics |
| LINURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 28 | 100% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 0 | 0% | 0 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 0 | 0% | 0 out of 20 |
| Plotted Points Statistics |
| MALAOXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 69% | 19% | 67% | 15% |
| Total | 22 | 69% | 19% | 67% | 15% |
| Sample Statistics |
| MALAOXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 88% | Spiked |
| Estimated Values | 6 | 24% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 25 | . | |
| False Negatives | 3 | 12% | 3 out of 25 |
| Not Spiked | 23 | . | |
| False Positives | 0 | 0% | 0 out of 23 |
| Plotted Points Statistics |
| MALATHION |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 110% | 33% | 103% | 17% |
| Total | 25 | 110% | 33% | 103% | 17% |
| Sample Statistics |
| MALATHION |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 93% | Spiked |
| Estimated Values | 6 | 22% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 2 | 7% | 2 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| Plotted Points Statistics |
| MCPA |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 18 | 124% | 29% | 123% | 28% |
| Total | 18 | 124% | 29% | 123% | 28% |
| Sample Statistics |
| MCPA |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 18 | 95% | Spiked |
| Estimated Values | 4 | 21% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 5% | Spiked |
| Spiked | 19 | . | |
| False Negatives | 0 | 0% | 0 out of 19 |
| Not Spiked | 29 | . | |
| False Positives | 4 | 14% | 4 out of 29 |
| Plotted Points Statistics |
| METALAXYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 29 | 100% | 12% | 100% | 9% |
| Total | 29 | 100% | 12% | 100% | 9% |
| Sample Statistics |
| METALAXYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 29 | 94% | Spiked |
| Estimated Values | 3 | 10% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 3% | Spiked |
| Spiked | 31 | . | |
| False Negatives | 1 | 3% | 1 out of 31 |
| Not Spiked | 17 | . | |
| False Positives | 1 | 6% | 1 out of 17 |
| Plotted Points Statistics |
| METCONAZOLE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 92% | 9% | 93% | 7% |
| Total | 21 | 92% | 9% | 93% | 7% |
| Sample Statistics |
| METCONAZOLE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 95% | Spiked |
| Estimated Values | 1 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 1 | 5% | 1 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| METHAMIDOPHOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 15 | 100% | 30% | 89% | 26% |
| Total | 15 | 100% | 30% | 89% | 26% |
| Sample Statistics |
| METHAMIDOPHOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 15 | 63% | Spiked |
| Estimated Values | 3 | 13% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 8% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 7 | 29% | 7 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 0 | 0% | 0 out of 24 |
| BQS OBSP Determination: |
| METHIDATHION (SUPRACIDE) |
| METHOD: 2437 , TESTIDCODE: 65088LCM60 |
| Open Data Set |
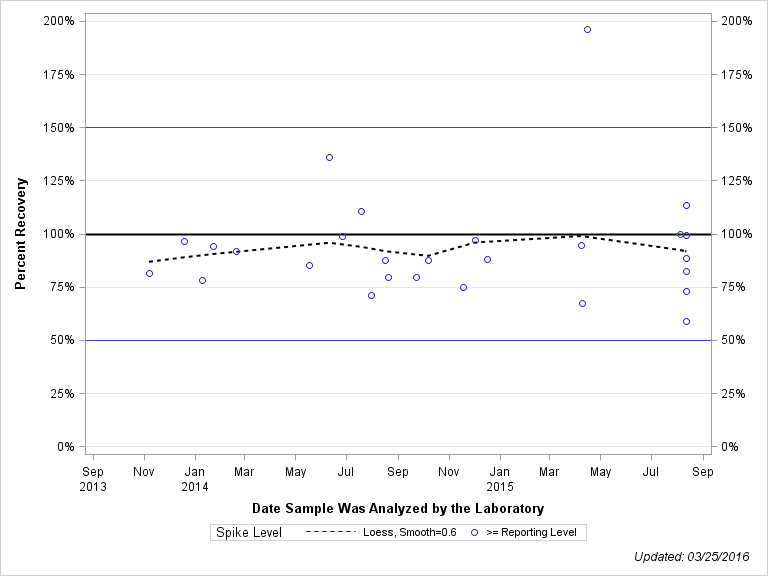
| Plotted Points Statistics |
| METHIDATHION (SUPRACIDE) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 27 | 93% | 26% | 88% | 14% |
| Total | 27 | 93% | 26% | 88% | 14% |
| Sample Statistics |
| METHIDATHION (SUPRACIDE) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 27 | 100% | Spiked |
| Estimated Values | 5 | 19% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 0 | 0% | 0 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| Plotted Points Statistics |
| METHOMYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 23 | 91% | 31% | 87% | 12% |
| Total | 23 | 91% | 31% | 87% | 12% |
| Sample Statistics |
| METHOMYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 23 | 96% | Spiked |
| Estimated Values | 5 | 21% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 0 | 0% | 0 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 3 | 13% | 3 out of 24 |
| Plotted Points Statistics |
| METHOMYL OXIME |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 15 | 88% | 21% | 85% | 24% |
| Total | 15 | 88% | 21% | 85% | 24% |
| Sample Statistics |
| METHOMYL OXIME |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 15 | 79% | Spiked |
| Estimated Values | 10 | 53% | Spiked |
| Deleted Values | 11 | 23% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 20 | . | |
| False Negatives | 4 | 21% | 4 out of 19 |
| Not Spiked | 28 | . | |
| False Positives | 0 | 0% | 0 out of 18 |
| Plotted Points Statistics |
| METHOXYFENOZIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 98% | 17% | 97% | 14% |
| Total | 25 | 98% | 17% | 97% | 14% |
| Sample Statistics |
| METHOXYFENOZIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 100% | Spiked |
| Estimated Values | 2 | 8% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 25 | . | |
| False Negatives | 0 | 0% | 0 out of 25 |
| Not Spiked | 23 | . | |
| False Positives | 0 | 0% | 0 out of 23 |
| Plotted Points Statistics |
| METHYL PARAOXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 19 | 110% | 35% | 114% | 32% |
| Total | 19 | 110% | 35% | 114% | 32% |
| Sample Statistics |
| METHYL PARAOXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 19 | 86% | Spiked |
| Estimated Values | 6 | 27% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 9% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 1 | 5% | 1 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| METOLACHLOR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 80% | 10% | 77% | 7% |
| Total | 26 | 80% | 10% | 77% | 7% |
| Sample Statistics |
| METOLACHLOR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 100% | Spiked |
| Estimated Values | 3 | 12% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 0 | 0% | 0 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 3 | 14% | 3 out of 22 |
| BQS OBSP Determination: |
| METOLACHLOR HYDROXY MORPHOLINONE |
| METHOD: 2437 , TESTIDCODE: 68649LCM60 |
| Open Data Set |

| Sample Statistics |
| METOLACHLOR HYDROXY MORPHOLINONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| BQS OBSP Determination: |
| METOLACHLOR OXANILIC ACID |
| METHOD: 2437 , TESTIDCODE: 68650LCM60 |
| Open Data Set |
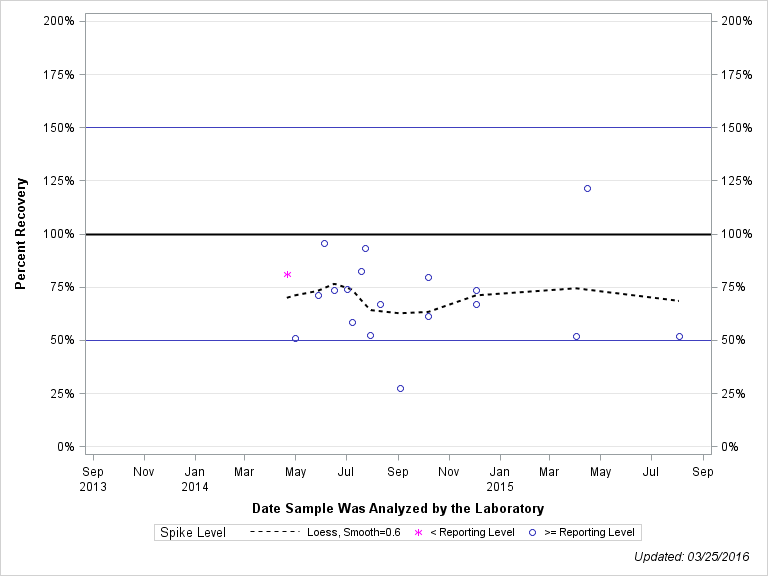
| Plotted Points Statistics |
| METOLACHLOR OXANILIC ACID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 1 | 81% | . | 81% | 0% |
| >= Reporting Level | 18 | 70% | 21% | 69% | 20% |
| Total | 19 | 70% | 21% | 71% | 21% |
| Sample Statistics |
| METOLACHLOR OXANILIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 19 | 90% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 21 | . | |
| False Negatives | 2 | 10% | 2 out of 21 |
| Not Spiked | 27 | . | |
| False Positives | 4 | 15% | 4 out of 27 |
| BQS OBSP Determination: |
| METOLACHLOR SULFONIC ACID |
| METHOD: 2437 , TESTIDCODE: 68651LCM60 |
| Open Data Set |
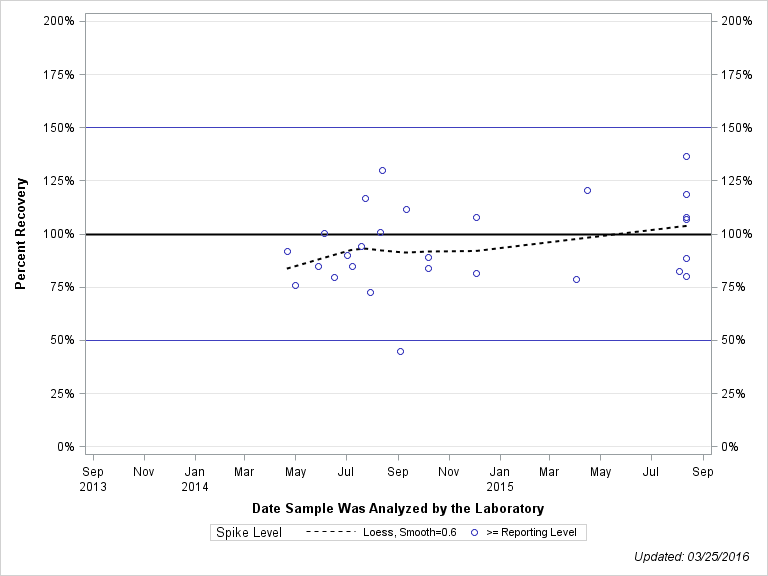
| Plotted Points Statistics |
| METOLACHLOR SULFONIC ACID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 27 | 95% | 20% | 90% | 20% |
| Total | 27 | 95% | 20% | 90% | 20% |
| Sample Statistics |
| METOLACHLOR SULFONIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 27 | 100% | Spiked |
| Estimated Values | 2 | 7% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 0 | 0% | 0 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| Plotted Points Statistics |
| METRIBUZIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 89% | 16% | 89% | 14% |
| Total | 26 | 89% | 16% | 89% | 14% |
| Sample Statistics |
| METRIBUZIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 0 | 0% | 0 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| METRIBUZIN-DESAMINO |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 6 | 81% | 9% | 84% | 13% |
| Total | 6 | 81% | 9% | 84% | 13% |
| Sample Statistics |
| METRIBUZIN-DESAMINO |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 6 | 100% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 6 | . | |
| False Negatives | 0 | 0% | 0 out of 6 |
| Not Spiked | 42 | . | |
| False Positives | 0 | 0% | 0 out of 42 |
| Plotted Points Statistics |
| METRIBUZIN DK |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 5 | 38% | 15% | 36% | 1% |
| Total | 5 | 38% | 15% | 36% | 1% |
| Sample Statistics |
| METRIBUZIN DK |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 5 | 83% | Spiked |
| Estimated Values | 2 | 33% | Spiked |
| Deleted Values | 3 | 6% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 8 | . | |
| False Negatives | 1 | 17% | 1 out of 6 |
| Not Spiked | 40 | . | |
| False Positives | 1 | 3% | 1 out of 39 |
| Plotted Points Statistics |
| MOLINATE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 28 | 126% | 30% | 127% | 29% |
| Total | 28 | 126% | 30% | 127% | 29% |
| Sample Statistics |
| MOLINATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 28 | 93% | Spiked |
| Estimated Values | 17 | 57% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 3% | Spiked |
| Spiked | 30 | . | |
| False Negatives | 1 | 3% | 1 out of 30 |
| Not Spiked | 18 | . | |
| False Positives | 0 | 0% | 0 out of 18 |
| Plotted Points Statistics |
| MYCLOBUTANIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 94% | 13% | 92% | 10% |
| Total | 21 | 94% | 13% | 92% | 10% |
| Sample Statistics |
| MYCLOBUTANIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 100% | Spiked |
| Estimated Values | 1 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 21 | . | |
| False Negatives | 0 | 0% | 0 out of 21 |
| Not Spiked | 27 | . | |
| False Positives | 0 | 0% | 0 out of 27 |
| BQS OBSP Determination: |
| N-(3,4-DICHLOROPHENYL)-N'-METHYLUREA |
| METHOD: 2437 , TESTIDCODE: 68231LCM60 |
| Open Data Set |
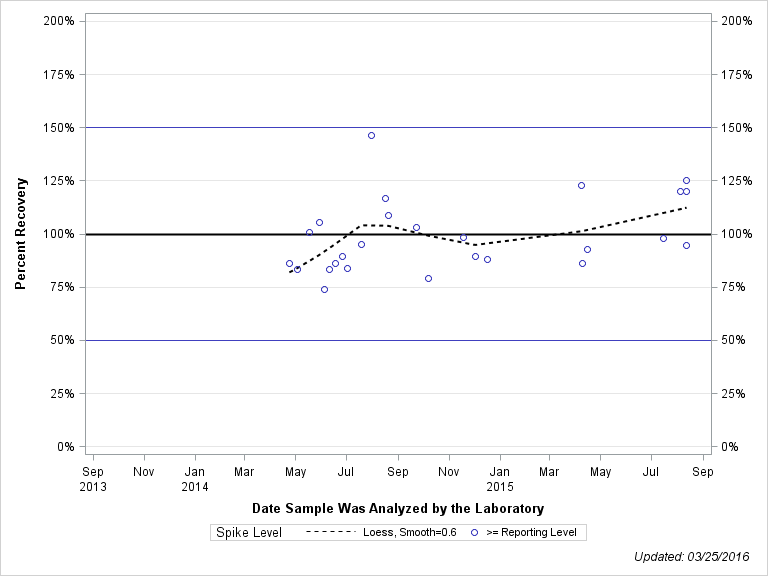
| Plotted Points Statistics |
| N-(3,4-DICHLOROPHENYL)-N'-METHYLUREA |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 99% | 17% | 95% | 17% |
| Total | 26 | 99% | 17% | 95% | 17% |
| Sample Statistics |
| N-(3,4-DICHLOROPHENYL)-N'-METHYLUREA |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 100% | Spiked |
| Estimated Values | 3 | 12% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 0 | 0% | 0 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 1 | 5% | 1 out of 22 |
| Plotted Points Statistics |
| NALED |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 14 | 138% | 78% | 128% | 114% |
| Total | 14 | 138% | 78% | 128% | 114% |
| Sample Statistics |
| NALED |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 14 | 93% | Spiked |
| Estimated Values | 14 | 93% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 15 | . | |
| False Negatives | 1 | 7% | 1 out of 15 |
| Not Spiked | 33 | . | |
| False Positives | 1 | 3% | 1 out of 33 |
| Plotted Points Statistics |
| NICOSULFURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 57% | 25% | 53% | 15% |
| Total | 21 | 57% | 25% | 53% | 15% |
| Sample Statistics |
| NICOSULFURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 95% | Spiked |
| Estimated Values | 7 | 32% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 1 | 5% | 1 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| NORFLURAZON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 102% | 12% | 103% | 11% |
| Total | 25 | 102% | 12% | 103% | 11% |
| Sample Statistics |
| NORFLURAZON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 96% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 1 | 4% | 1 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| NOVALURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 14 | 70% | 18% | 72% | 16% |
| Total | 14 | 70% | 18% | 72% | 16% |
| Sample Statistics |
| NOVALURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 14 | 82% | Spiked |
| Estimated Values | 14 | 82% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 17 | . | |
| False Negatives | 3 | 18% | 3 out of 17 |
| Not Spiked | 31 | . | |
| False Positives | 0 | 0% | 0 out of 31 |
| BQS OBSP Determination: |
| O-ETHYL-O-METHYL-S-PROPYLPHOPHOROTHIOATE |
| METHOD: 2437 , TESTIDCODE: 68597LCM60 |
| Open Data Set |

| Sample Statistics |
| O-ETHYL-O-METHYL-S-PROPYLPHOPHOROTHIOATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 1 | 2% | 1 out of 48 |
| BQS OBSP Determination: |
| O-ETHYL-S-METHYL-S-PROPYL PHOSPHORODITHIOATE |
| METHOD: 2437 , TESTIDCODE: 68657LCM60 |
| Open Data Set |

| Sample Statistics |
| O-ETHYL-S-METHYL-S-PROPYL PHOSPHORODITHIOATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| BQS OBSP Determination: |
| O-ETHYL-S-PROPYLPHOSPHOROTHIOATE |
| METHOD: 2437 , TESTIDCODE: 68658LCM60 |
| Open Data Set |

| Sample Statistics |
| O-ETHYL-S-PROPYLPHOSPHOROTHIOATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| ORTHOSULFAMURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 10 | 32% | 15% | 35% | 7% |
| Total | 10 | 32% | 15% | 35% | 7% |
| Sample Statistics |
| ORTHOSULFAMURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 10 | 59% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 17 | . | |
| False Negatives | 7 | 41% | 7 out of 17 |
| Not Spiked | 31 | . | |
| False Positives | 0 | 0% | 0 out of 31 |
| Plotted Points Statistics |
| ORYZALIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 23 | 109% | 24% | 103% | 18% |
| Total | 23 | 109% | 24% | 103% | 18% |
| Sample Statistics |
| ORYZALIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 23 | 100% | Spiked |
| Estimated Values | 3 | 13% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 23 | . | |
| False Negatives | 0 | 0% | 0 out of 23 |
| Not Spiked | 25 | . | |
| False Positives | 0 | 0% | 0 out of 25 |
| Plotted Points Statistics |
| OXAMYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 19 | 74% | 20% | 76% | 21% |
| Total | 19 | 74% | 20% | 76% | 21% |
| Sample Statistics |
| OXAMYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 19 | 86% | Spiked |
| Estimated Values | 3 | 14% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 3 | 14% | 3 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 1 | 4% | 1 out of 26 |
| Plotted Points Statistics |
| OXAMYL OXIME |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 1 | 18% | . | 18% | 0% |
| Total | 1 | 18% | . | 18% | 0% |
| Sample Statistics |
| OXAMYL OXIME |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 1 | 6% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 17 | . | |
| False Negatives | 16 | 94% | 16 out of 17 |
| Not Spiked | 31 | . | |
| False Positives | 4 | 13% | 4 out of 31 |
| Plotted Points Statistics |
| OXYFLUORFEN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| < Reporting Level | 2 | 50% | 26% | 50% | 27% |
| >= Reporting Level | 16 | 92% | 21% | 91% | 10% |
| Total | 18 | 88% | 25% | 91% | 16% |
| Sample Statistics |
| OXYFLUORFEN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 18 | 90% | Spiked |
| Estimated Values | 18 | 90% | Spiked |
| Deleted Values | 2 | 4% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 10% | Spiked |
| Spiked | 21 | . | |
| False Negatives | 0 | 0% | 0 out of 20 |
| Not Spiked | 27 | . | |
| False Positives | 4 | 15% | 4 out of 26 |
| Plotted Points Statistics |
| PARAOXON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 108% | 30% | 108% | 22% |
| Total | 21 | 108% | 30% | 108% | 22% |
| Sample Statistics |
| PARAOXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 100% | Spiked |
| Estimated Values | 8 | 38% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 21 | . | |
| False Negatives | 0 | 0% | 0 out of 21 |
| Not Spiked | 27 | . | |
| False Positives | 0 | 0% | 0 out of 27 |
| Sample Statistics |
| PARATHION-METHYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | 0% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 28 | 70% | Spiked + Not Spiked |
| Spiked, Censored | 7 | 100% | Spiked |
| Spiked | 23 | . | |
| False Negatives | 0 | 0% | 0 out of 7 |
| Not Spiked | 17 | . | |
| False Positives | 3 | 60% | 3 out of 5 |
| Plotted Points Statistics |
| PENDIMETHALIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 14 | 92% | 12% | 93% | 11% |
| Total | 14 | 92% | 12% | 93% | 11% |
| Sample Statistics |
| PENDIMETHALIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 14 | 54% | Spiked |
| Estimated Values | 4 | 15% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 5 | 19% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 7 | 27% | 7 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 0 | 0% | 0 out of 22 |
| Plotted Points Statistics |
| PHORATE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 20 | 86% | 28% | 83% | 34% |
| Total | 20 | 86% | 28% | 83% | 34% |
| Sample Statistics |
| PHORATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 20 | 74% | Spiked |
| Estimated Values | 2 | 7% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 7% | Spiked |
| Spiked | 27 | . | |
| False Negatives | 5 | 19% | 5 out of 27 |
| Not Spiked | 21 | . | |
| False Positives | 0 | 0% | 0 out of 21 |
| Plotted Points Statistics |
| PHORATE OXON SULFOXIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 5 | 96% | 12% | 91% | 13% |
| Total | 5 | 96% | 12% | 91% | 13% |
| Sample Statistics |
| PHORATE OXON SULFOXIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 5 | 83% | Spiked |
| Estimated Values | 2 | 33% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 6 | . | |
| False Negatives | 1 | 17% | 1 out of 6 |
| Not Spiked | 42 | . | |
| False Positives | 4 | 10% | 4 out of 42 |
| Plotted Points Statistics |
| PHORATE OXYGEN ANALOG |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 10 | 83% | 28% | 79% | 13% |
| Total | 10 | 83% | 28% | 79% | 13% |
| Sample Statistics |
| PHORATE OXYGEN ANALOG |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 10 | 43% | Spiked |
| Estimated Values | 3 | 13% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 8 | 35% | Spiked |
| Spiked | 23 | . | |
| False Negatives | 5 | 22% | 5 out of 23 |
| Not Spiked | 25 | . | |
| False Positives | 1 | 4% | 1 out of 25 |
| BQS OBSP Determination: |
| PHORATE OXYGEN ANALOG SULFONE |
| METHOD: 2437 , TESTIDCODE: 68670LCM60 |
| Open Data Set |
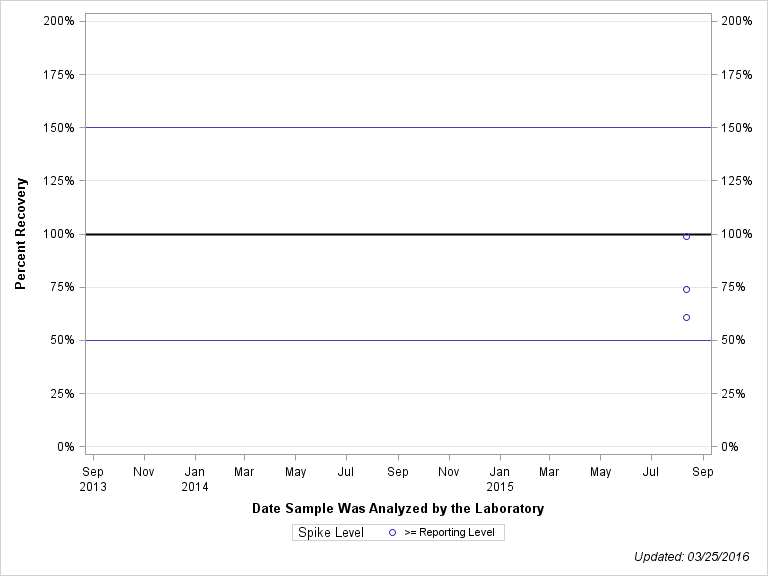
| Plotted Points Statistics |
| PHORATE OXYGEN ANALOG SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 3 | 78% | 19% | 74% | 28% |
| Total | 3 | 78% | 19% | 74% | 28% |
| Sample Statistics |
| PHORATE OXYGEN ANALOG SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 3 | 100% | Spiked |
| Estimated Values | 3 | 100% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 3 | . | |
| False Negatives | 0 | 0% | 0 out of 3 |
| Not Spiked | 45 | . | |
| False Positives | 0 | 0% | 0 out of 45 |
| Plotted Points Statistics |
| PHORATE SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 16 | 102% | 47% | 90% | 43% |
| Total | 16 | 102% | 47% | 90% | 43% |
| Sample Statistics |
| PHORATE SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 16 | 89% | Spiked |
| Estimated Values | 3 | 17% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 18 | . | |
| False Negatives | 2 | 11% | 2 out of 18 |
| Not Spiked | 30 | . | |
| False Positives | 1 | 3% | 1 out of 30 |
| Plotted Points Statistics |
| PHORATE SULFOXIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 20 | 117% | 47% | 103% | 34% |
| Total | 20 | 117% | 47% | 103% | 34% |
| Sample Statistics |
| PHORATE SULFOXIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 20 | 100% | Spiked |
| Estimated Values | 1 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 20 | . | |
| False Negatives | 0 | 0% | 0 out of 20 |
| Not Spiked | 28 | . | |
| False Positives | 15 | 54% | 15 out of 28 |
| Sample Statistics |
| PHOSMET (IMIDAN) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 40 | 100% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 20 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 20 | . | |
| False Positives | 0 | . | 0 out of 0 |
| Plotted Points Statistics |
| PHTHALAZINONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 5 | 255% | 90% | 252% | 78% |
| Total | 5 | 255% | 90% | 252% | 78% |
| Sample Statistics |
| PHTHALAZINONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 5 | 63% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 8 | . | |
| False Negatives | 3 | 38% | 3 out of 8 |
| Not Spiked | 40 | . | |
| False Positives | 14 | 35% | 14 out of 40 |
| Plotted Points Statistics |
| PIPERONYL BUTOXIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 10 | 60% | 20% | 60% | 13% |
| Total | 10 | 60% | 20% | 60% | 13% |
| Sample Statistics |
| PIPERONYL BUTOXIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 10 | 56% | Spiked |
| Estimated Values | 2 | 11% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 7 | 39% | Spiked |
| Spiked | 18 | . | |
| False Negatives | 1 | 6% | 1 out of 18 |
| Not Spiked | 30 | . | |
| False Positives | 1 | 3% | 1 out of 30 |
| Plotted Points Statistics |
| PROFENOFOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 18 | 97% | 25% | 105% | 30% |
| Total | 18 | 97% | 25% | 105% | 30% |
| Sample Statistics |
| PROFENOFOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 18 | 100% | Spiked |
| Estimated Values | 3 | 17% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 18 | . | |
| False Negatives | 0 | 0% | 0 out of 18 |
| Not Spiked | 30 | . | |
| False Positives | 0 | 0% | 0 out of 30 |
| Plotted Points Statistics |
| PROMETON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 94% | 11% | 97% | 8% |
| Total | 25 | 94% | 11% | 97% | 8% |
| Sample Statistics |
| PROMETON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 96% | Spiked |
| Estimated Values | 2 | 8% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 1 | 4% | 1 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 6 | 27% | 6 out of 22 |
| Plotted Points Statistics |
| PROMETRYN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 80% | 28% | 83% | 9% |
| Total | 21 | 80% | 28% | 83% | 9% |
| Sample Statistics |
| PROMETRYN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 91% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 23 | . | |
| False Negatives | 2 | 9% | 2 out of 23 |
| Not Spiked | 25 | . | |
| False Positives | 3 | 12% | 3 out of 25 |
| Plotted Points Statistics |
| PROPANIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 30 | 106% | 14% | 106% | 12% |
| Total | 30 | 106% | 14% | 106% | 12% |
| Sample Statistics |
| PROPANIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 30 | 100% | Spiked |
| Estimated Values | 1 | 3% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 30 | . | |
| False Negatives | 0 | 0% | 0 out of 30 |
| Not Spiked | 18 | . | |
| False Positives | 0 | 0% | 0 out of 18 |
| Plotted Points Statistics |
| PROPARGITE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 78% | 24% | 81% | 18% |
| Total | 25 | 78% | 24% | 81% | 18% |
| Sample Statistics |
| PROPARGITE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 86% | Spiked |
| Estimated Values | 5 | 17% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 3% | Spiked |
| Spiked | 29 | . | |
| False Negatives | 3 | 10% | 3 out of 29 |
| Not Spiked | 19 | . | |
| False Positives | 0 | 0% | 0 out of 19 |
| Plotted Points Statistics |
| PROPAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 92% | 18% | 90% | 14% |
| Total | 22 | 92% | 18% | 90% | 14% |
| Sample Statistics |
| PROPAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 96% | Spiked |
| Estimated Values | 2 | 9% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 23 | . | |
| False Negatives | 1 | 4% | 1 out of 23 |
| Not Spiked | 25 | . | |
| False Positives | 7 | 28% | 7 out of 25 |
| Plotted Points Statistics |
| PROPICONAZOLE (TILT) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 28 | 84% | 13% | 85% | 13% |
| Total | 28 | 84% | 13% | 85% | 13% |
| Sample Statistics |
| PROPICONAZOLE (TILT) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 28 | 90% | Spiked |
| Estimated Values | 2 | 6% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 31 | . | |
| False Negatives | 3 | 10% | 3 out of 31 |
| Not Spiked | 17 | . | |
| False Positives | 0 | 0% | 0 out of 17 |
| Plotted Points Statistics |
| PROPOXUR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 23 | 97% | 24% | 93% | 17% |
| Total | 23 | 97% | 24% | 93% | 17% |
| Sample Statistics |
| PROPOXUR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 23 | 100% | Spiked |
| Estimated Values | 3 | 13% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 23 | . | |
| False Negatives | 0 | 0% | 0 out of 23 |
| Not Spiked | 25 | . | |
| False Positives | 0 | 0% | 0 out of 25 |
| Plotted Points Statistics |
| PROPYZAMIDE (PRONAMIDE) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 29 | 110% | 23% | 109% | 17% |
| Total | 29 | 110% | 23% | 109% | 17% |
| Sample Statistics |
| PROPYZAMIDE (PRONAMIDE) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 29 | 97% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 3% | Spiked |
| Spiked | 30 | . | |
| False Negatives | 0 | 0% | 0 out of 30 |
| Not Spiked | 18 | . | |
| False Positives | 0 | 0% | 0 out of 18 |
| Plotted Points Statistics |
| PROSULFURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 20 | 70% | 13% | 68% | 9% |
| Total | 20 | 70% | 13% | 68% | 9% |
| Sample Statistics |
| PROSULFURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 20 | 87% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 23 | . | |
| False Negatives | 3 | 13% | 3 out of 23 |
| Not Spiked | 25 | . | |
| False Positives | 0 | 0% | 0 out of 25 |
| Sample Statistics |
| PYMETROZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 40 | 83% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 17 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 31 | . | |
| False Positives | 0 | 0% | 0 out of 8 |
| Plotted Points Statistics |
| PYRACLOSTROBIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 18 | 79% | 11% | 82% | 11% |
| Total | 18 | 79% | 11% | 82% | 11% |
| Sample Statistics |
| PYRACLOSTROBIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 18 | 90% | Spiked |
| Estimated Values | 1 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 20 | . | |
| False Negatives | 2 | 10% | 2 out of 20 |
| Not Spiked | 28 | . | |
| False Positives | 0 | 0% | 0 out of 28 |
| Plotted Points Statistics |
| PYRIDABEN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 16 | 62% | 20% | 64% | 12% |
| Total | 16 | 62% | 20% | 64% | 12% |
| Sample Statistics |
| PYRIDABEN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 16 | 94% | Spiked |
| Estimated Values | 2 | 12% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 17 | . | |
| False Negatives | 1 | 6% | 1 out of 17 |
| Not Spiked | 31 | . | |
| False Positives | 0 | 0% | 0 out of 31 |
| Plotted Points Statistics |
| PYRIPROXYFEN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 19 | 88% | 13% | 86% | 14% |
| Total | 19 | 88% | 13% | 86% | 14% |
| Sample Statistics |
| PYRIPROXYFEN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 19 | 100% | Spiked |
| Estimated Values | 8 | 42% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 19 | . | |
| False Negatives | 0 | 0% | 0 out of 19 |
| Not Spiked | 29 | . | |
| False Positives | 0 | 0% | 0 out of 29 |
| BQS OBSP Determination: |
| SEC-ACETOCHLOR OXANILIC ACID |
| METHOD: 2437 , TESTIDCODE: 68684LCM60 |
| Open Data Set |
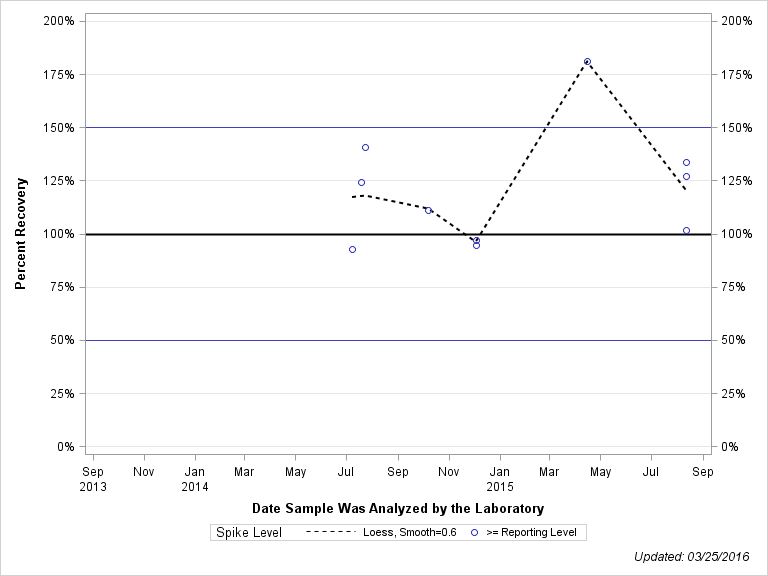
| Plotted Points Statistics |
| SEC-ACETOCHLOR OXANILIC ACID |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 10 | 120% | 27% | 118% | 27% |
| Total | 10 | 120% | 27% | 118% | 27% |
| Sample Statistics |
| SEC-ACETOCHLOR OXANILIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 10 | 100% | Spiked |
| Estimated Values | 1 | 10% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 10 | . | |
| False Negatives | 0 | 0% | 0 out of 10 |
| Not Spiked | 38 | . | |
| False Positives | 0 | 0% | 0 out of 38 |
| BQS OBSP Determination: |
| SEC-ALACHLOR OXANILIC ACID |
| METHOD: 2437 , TESTIDCODE: 68685LCM60 |
| Open Data Set |

| Sample Statistics |
| SEC-ALACHLOR OXANILIC ACID |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| SIDURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 24 | 96% | 19% | 95% | 24% |
| Total | 24 | 96% | 19% | 95% | 24% |
| Sample Statistics |
| SIDURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 24 | 96% | Spiked |
| Estimated Values | 2 | 8% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 25 | . | |
| False Negatives | 1 | 4% | 1 out of 25 |
| Not Spiked | 23 | . | |
| False Positives | 0 | 0% | 0 out of 23 |
| Plotted Points Statistics |
| SIMAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 26 | 102% | 23% | 95% | 18% |
| Total | 26 | 102% | 23% | 95% | 18% |
| Sample Statistics |
| SIMAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 26 | 100% | Spiked |
| Estimated Values | 4 | 15% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 0 | 0% | 0 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 1 | 5% | 1 out of 22 |
| Plotted Points Statistics |
| SULFENTRAZONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 101% | 27% | 109% | 30% |
| Total | 21 | 101% | 27% | 109% | 30% |
| Sample Statistics |
| SULFENTRAZONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 91% | Spiked |
| Estimated Values | 2 | 9% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 2 | 9% | Spiked |
| Spiked | 23 | . | |
| False Negatives | 0 | 0% | 0 out of 23 |
| Not Spiked | 25 | . | |
| False Positives | 0 | 0% | 0 out of 25 |
| Plotted Points Statistics |
| SULFOMETURON-METHYL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 20 | 94% | 29% | 100% | 21% |
| Total | 20 | 94% | 29% | 100% | 21% |
| Sample Statistics |
| SULFOMETURON-METHYL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 20 | 95% | Spiked |
| Estimated Values | 7 | 33% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 21 | . | |
| False Negatives | 1 | 5% | 1 out of 21 |
| Not Spiked | 27 | . | |
| False Positives | 0 | 0% | 0 out of 27 |
| Plotted Points Statistics |
| SULFOSULFURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 15 | 134% | 31% | 122% | 21% |
| Total | 15 | 134% | 31% | 122% | 21% |
| Sample Statistics |
| SULFOSULFURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 15 | 83% | Spiked |
| Estimated Values | 1 | 6% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 18 | . | |
| False Negatives | 3 | 17% | 3 out of 18 |
| Not Spiked | 30 | . | |
| False Positives | 0 | 0% | 0 out of 30 |
| BQS OBSP Determination: |
| SULFOSULFURON ETHYL SULFONE |
| METHOD: 2437 , TESTIDCODE: 68690LCM60 |
| Open Data Set |

| Sample Statistics |
| SULFOSULFURON ETHYL SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Sample Statistics |
| TCPSA, ETHYL ESTER |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| TEBUCONAZOLE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 91% | 13% | 88% | 11% |
| Total | 21 | 91% | 13% | 88% | 11% |
| Sample Statistics |
| TEBUCONAZOLE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 91% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 23 | . | |
| False Negatives | 1 | 4% | 1 out of 23 |
| Not Spiked | 25 | . | |
| False Positives | 0 | 0% | 0 out of 25 |
| Plotted Points Statistics |
| TEBUFENOZIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 19 | 76% | 20% | 68% | 17% |
| Total | 19 | 76% | 20% | 68% | 17% |
| Sample Statistics |
| TEBUFENOZIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 19 | 100% | Spiked |
| Estimated Values | 3 | 16% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 19 | . | |
| False Negatives | 0 | 0% | 0 out of 19 |
| Not Spiked | 29 | . | |
| False Positives | 0 | 0% | 0 out of 29 |
| Sample Statistics |
| TEBUPIRIMFOS OXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| BQS OBSP Determination: |
| TEBUPIRIMPHOS (PHOSTEBUPIRIM) |
| METHOD: 2437 , TESTIDCODE: 68693LCM60 |
| Open Data Set |
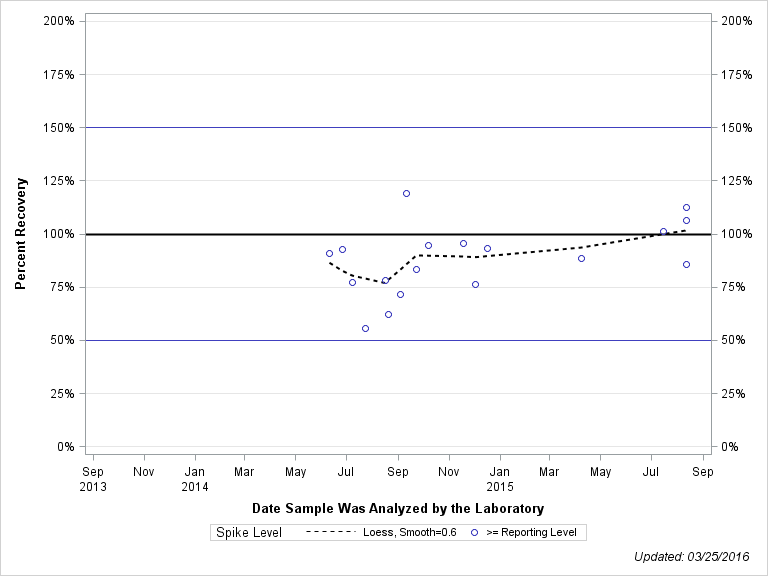
| Plotted Points Statistics |
| TEBUPIRIMPHOS (PHOSTEBUPIRIM) |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 18 | 88% | 16% | 90% | 13% |
| Total | 18 | 88% | 16% | 90% | 13% |
| Sample Statistics |
| TEBUPIRIMPHOS (PHOSTEBUPIRIM) |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 18 | 95% | Spiked |
| Estimated Values | 5 | 26% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 19 | . | |
| False Negatives | 1 | 5% | 1 out of 19 |
| Not Spiked | 29 | . | |
| False Positives | 0 | 0% | 0 out of 29 |
| Plotted Points Statistics |
| TEBUTHIURON |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 32 | 96% | 17% | 96% | 11% |
| Total | 32 | 96% | 17% | 96% | 11% |
| Sample Statistics |
| TEBUTHIURON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 32 | 100% | Spiked |
| Estimated Values | 4 | 13% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 32 | . | |
| False Negatives | 0 | 0% | 0 out of 32 |
| Not Spiked | 16 | . | |
| False Positives | 0 | 0% | 0 out of 16 |
| Sample Statistics |
| TEBUTHIURON TP 104 |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| TEBUTHIURON TP EL108 |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 15 | 92% | 16% | 95% | 15% |
| Total | 15 | 92% | 16% | 95% | 15% |
| Sample Statistics |
| TEBUTHIURON TP EL108 |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 15 | 94% | Spiked |
| Estimated Values | 0 | 0% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 16 | . | |
| False Negatives | 1 | 6% | 1 out of 16 |
| Not Spiked | 32 | . | |
| False Positives | 0 | 0% | 0 out of 32 |
| Sample Statistics |
| TEBUTHIURON TP EL109 |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| BQS OBSP Determination: |
| TEBUTHIURON TRANSFORMATION PRODUCT 106 |
| METHOD: 2437 , TESTIDCODE: 68714LCM60 |
| Open Data Set |

| Sample Statistics |
| TEBUTHIURON TRANSFORMATION PRODUCT 106 |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| TERBACIL |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 89% | 20% | 96% | 23% |
| Total | 22 | 89% | 20% | 96% | 23% |
| Sample Statistics |
| TERBACIL |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 100% | Spiked |
| Estimated Values | 1 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 0 | 0% | 0 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 4 | 15% | 4 out of 26 |
| Plotted Points Statistics |
| TERBUFOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 25 | 69% | 21% | 70% | 17% |
| Total | 25 | 69% | 21% | 70% | 17% |
| Sample Statistics |
| TERBUFOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 25 | 89% | Spiked |
| Estimated Values | 4 | 14% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 28 | . | |
| False Negatives | 3 | 11% | 3 out of 28 |
| Not Spiked | 20 | . | |
| False Positives | 0 | 0% | 0 out of 20 |
| Sample Statistics |
| TERBUFOS OXON |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Sample Statistics |
| TERBUFOS OXON SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 1 | 2% | 1 out of 48 |
| Sample Statistics |
| TERBUFOS OXON SULFOXIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 0 | . | Spiked |
| Estimated Values | 0 | . | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | . | Spiked |
| Spiked | 0 | . | |
| False Negatives | 0 | . | 0 out of 0 |
| Not Spiked | 48 | . | |
| False Positives | 0 | 0% | 0 out of 48 |
| Plotted Points Statistics |
| TERBUFOS SULFONE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 21 | 106% | 32% | 106% | 20% |
| Total | 21 | 106% | 32% | 106% | 20% |
| Sample Statistics |
| TERBUFOS SULFONE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 21 | 100% | Spiked |
| Estimated Values | 1 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 21 | . | |
| False Negatives | 0 | 0% | 0 out of 21 |
| Not Spiked | 27 | . | |
| False Positives | 0 | 0% | 0 out of 27 |
| Plotted Points Statistics |
| TERBUFOS SULFOXIDE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 19 | 113% | 38% | 115% | 44% |
| Total | 19 | 113% | 38% | 115% | 44% |
| Sample Statistics |
| TERBUFOS SULFOXIDE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 19 | 100% | Spiked |
| Estimated Values | 2 | 11% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 19 | . | |
| False Negatives | 0 | 0% | 0 out of 19 |
| Not Spiked | 29 | . | |
| False Positives | 16 | 55% | 16 out of 29 |
| Plotted Points Statistics |
| TERBUTHYLAZINE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 93% | 14% | 93% | 10% |
| Total | 22 | 93% | 14% | 93% | 10% |
| Sample Statistics |
| TERBUTHYLAZINE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 96% | Spiked |
| Estimated Values | 1 | 4% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 23 | . | |
| False Negatives | 1 | 4% | 1 out of 23 |
| Not Spiked | 25 | . | |
| False Positives | 2 | 8% | 2 out of 25 |
| Plotted Points Statistics |
| TETRACONAZOLE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 20 | 85% | 16% | 83% | 14% |
| Total | 20 | 85% | 16% | 83% | 14% |
| Sample Statistics |
| TETRACONAZOLE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 20 | 91% | Spiked |
| Estimated Values | 1 | 5% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 5% | Spiked |
| Spiked | 22 | . | |
| False Negatives | 1 | 5% | 1 out of 22 |
| Not Spiked | 26 | . | |
| False Positives | 0 | 0% | 0 out of 26 |
| Plotted Points Statistics |
| THIOBENCARB |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 29 | 79% | 16% | 74% | 13% |
| Total | 29 | 79% | 16% | 74% | 13% |
| Sample Statistics |
| THIOBENCARB |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 29 | 97% | Spiked |
| Estimated Values | 4 | 13% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 30 | . | |
| False Negatives | 1 | 3% | 1 out of 30 |
| Not Spiked | 18 | . | |
| False Positives | 0 | 0% | 0 out of 18 |
| Plotted Points Statistics |
| TRANS-PERMETHRIN |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 18 | 54% | 14% | 51% | 12% |
| Total | 18 | 54% | 14% | 51% | 12% |
| Sample Statistics |
| TRANS-PERMETHRIN |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 18 | 95% | Spiked |
| Estimated Values | 7 | 37% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 19 | . | |
| False Negatives | 1 | 5% | 1 out of 19 |
| Not Spiked | 29 | . | |
| False Positives | 2 | 7% | 2 out of 29 |
| Plotted Points Statistics |
| TRI-ALLATE |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 24 | 82% | 16% | 81% | 13% |
| Total | 24 | 82% | 16% | 81% | 13% |
| Sample Statistics |
| TRI-ALLATE |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 24 | 100% | Spiked |
| Estimated Values | 9 | 38% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 24 | . | |
| False Negatives | 0 | 0% | 0 out of 24 |
| Not Spiked | 24 | . | |
| False Positives | 0 | 0% | 0 out of 24 |
| Plotted Points Statistics |
| TRIBUFOS |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 22 | 79% | 15% | 80% | 15% |
| Total | 22 | 79% | 15% | 80% | 15% |
| Sample Statistics |
| TRIBUFOS |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 22 | 85% | Spiked |
| Estimated Values | 4 | 15% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 1 | 4% | Spiked |
| Spiked | 26 | . | |
| False Negatives | 3 | 12% | 3 out of 26 |
| Not Spiked | 22 | . | |
| False Positives | 1 | 5% | 1 out of 22 |
| Plotted Points Statistics |
| TRICLOPYR |
| Spike Level | N | Mean | Std-Dev. | Median | F_Pseudo |
|---|---|---|---|---|---|
| >= Reporting Level | 19 | 83% | 19% | 86% | 19% |
| Total | 19 | 83% | 19% | 86% | 19% |
| Sample Statistics |
| TRICLOPYR |
| Characteristic | N | % | % Basis |
|---|---|---|---|
| Plotted | 19 | 100% | Spiked |
| Estimated Values | 5 | 26% | Spiked |
| Deleted Values | 0 | 0% | Spiked + Not Spiked |
| Spiked, Censored | 0 | 0% | Spiked |
| Spiked | 19 | . | |
| False Negatives | 0 | 0% | 0 out of 19 |
| Not Spiked | 29 | . | |
| False Positives | 4 | 14% | 4 out of 29 |